 Your new post is loading...
 Your new post is loading...
Patients do not access physicians at random but rather via naturally emerging
networks of patient flows between them. As retirements, mass quarantines and absence due to sickness during pandemics, or other shocks thin out these networks, the system might be pushed closer to a tipping point where...
Analysis: With experts concerned over rising UK case rates, where are we also with deaths, hospital admissions, long Covid and the economy?
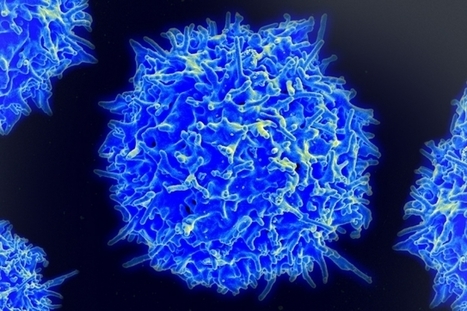
Researchers at MIT have developed a synthetic gene circuit that triggers the body’s immune system to attack cancers when it detects signs of the disease.
The circuit, which will only activate a therapeutic response when it detects two specific cancer markers, is described in a paper published today in the journal Cell.
Immunotherapy is widely seen as having considerable potential in the fight against a range of cancers. The approach has been demonstrated successfully in several recent clinical trials, according to Timothy Lu, associate professor of biological engineering and of electrical engineering and computer science at MIT.
“There has been a lot of clinical data recently suggesting that if you can stimulate the immune system in the right way you can get it to recognize cancer,” says Lu, who is head of the Synthetic Biology Group in MIT’s Research Laboratory of Electronics. “Some of the best examples of this are what are called checkpoint inhibitors, where essentially cancers put up stop signs [that prevent] T-cells from killing them. There are antibodies that have been developed now that basically block those inhibitory signals and allow the immune system to act against the cancers.”
However, despite this success, the use of immunotherapy remains limited by the scarcity of tumor-specific antigens — substances that can trigger an immune system response to a particular type of cancer. The toxicity of some therapies, when delivered as a systemic treatment to the whole body, for example, is another obstacle.
What’s more, the treatments are not successful in all cases. Indeed, even in some of the most successful tests, only 30-40 percent of patients will respond to a given therapy, Lu says.
As a result, there is now a push to develop combination therapies, in which different but complementary treatments are used to boost the immune response. So, for example, if one type of immunotherapy is used to knock out an inhibitory signal produced by a cancer, and the tumor responds by upregulating a second signal, an additional therapy could then be used to target this one as well, Lu says.
“Our belief is that there is a need to develop much more specific, targeted immunotherapies that work locally at the tumor site, rather than trying to treat the entire body systemically,” he says. “Secondly, we want to produce multiple immunotherapies from a single package, and therefore be able to stimulate the immune system in multiple different ways.”
To do this, Lu and a team including MIT postdocs Lior Nissim and Ming-Ru Wu, have built a gene circuit encoded in DNA designed to distinguish cancer cells from noncancer cells.
The circuit, which can be customized to respond to different types of tumor, is based on the simple AND gates used in electronics. Such AND gates will only switch on a circuit when two inputs are present.
Cancer cells differ from normal cells in the profile of their gene expression. So the researchers developed synthetic promoters — DNA sequences designed to initiate gene expression but only in cancer cells.
The circuit is delivered to cells in the affected area of the body using a virus. The synthetic promotors are then designed to bind to certain proteins that are active in tumor cells, causing the promoters to turn on.
“Only when two of these cancer promoters are activated, does the circuit itself switch on,” Lu says.
This allows the circuit to target tumors more accurately than existing therapies, as it requires two cancer-specific signals to be present before it will respond.
Once activated, the circuit expresses proteins designed to direct the immune system to target the tumor cells, including surface T cell engagers, which direct T cells to kill the cells. The circuit also expresses a checkpoint inhibitor designed to lift the brakes on T cell activity.
When the researchers tested the circuit in vitro, they found that it was able to detect ovarian cancer cells from amongst other noncancerous ovarian cells and other cell types.
They then tested the circuit in mice implanted with ovarian cancer cells, and demonstrated that it could trigger T cells to seek out and kill the cancer cells without harming other cells around them.
Finally, the researchers showed that the circuit could be readily converted to target other cancer cells.
“We identified other promoters that were selective for breast cancer, and when these were encoded into the circuit, it would target breast cancer cells over other types of cell,” Lu says.
Ultimately, they hope they will also be able to use the system to target other diseases, such as rheumatoid arthritis, inflammatory bowel disease, and other autoimmune diseases.
This advance will open up a new front against cancer, says Martin Fussenegger, a professor of biotechnology and bioengineering at ETH Zurich in Switzerland, who was not involved in the research.
“First author Lior Nissim, who pioneered the very first genetic circuit targeting tumor cells, has now teamed up with Timothy Lu to design RNA-based immunomodulatory gene circuits that take cancer immunotherapy to a new level,” Fussenegger says. “The design of this highly complex tumor-killing gene circuit was made possible by meticulous optimization and integration of several components that target and program tumor cells to become a specific prey for the immune system — this is very smart technology.”
The researchers now plan to test the circuit more fully in a range of cancer models. They are also aiming to develop a delivery system for the circuit, which would be both flexible and simple to manufacture and use.
Via Gerd Moe-Behrens
It will enter clinical trials to prevent and treat the infection next year.
The majority of human emerging infectious diseases are zoonotic, with viruses that originate in wild mammals of particular concern (for example, HIV, Ebola and SARS). Understanding patterns of viral diversity in wildlife and determinants of successful cross-species transmission, or spillover, are therefore key goals for pandemic surveillance programs. However, few analytical tools exist to identify which host species are likely to harbour the next human virus, or which viruses can cross species boundaries. Here we conduct a comprehensive analysis of mammalian host–virus relationships and show that both the total number of viruses that infect a given species and the proportion likely to be zoonotic are predictable. After controlling for research effort, the proportion of zoonotic viruses per species is predicted by phylogenetic relatedness to humans, host taxonomy and human population within a species range—which may reflect human–wildlife contact. We demonstrate that bats harbour a significantly higher proportion of zoonotic viruses than all other mammalian orders. We also identify the taxa and geographic regions with the largest estimated number of ‘missing viruses’ and ‘missing zoonoses’ and therefore of highest value for future surveillance. We then show that phylogenetic host breadth and other viral traits are significant predictors of zoonotic potential, providing a novel framework to assess if a newly discovered mammalian virus could infect people.
Via Ed Rybicki
One hundred and ten Zika virus genomes from ten countries and territories involved in the Zika virus epidemic reveal rapid expansion of the epidemic within Brazil and multiple introductions to other regions. Zika virus evolution and spread in the Americas
Hayden C. Metsky, et al. Nature 546, 411–415 (15 June 2017) doi:10.1038/nature22402
Via Complexity Digest
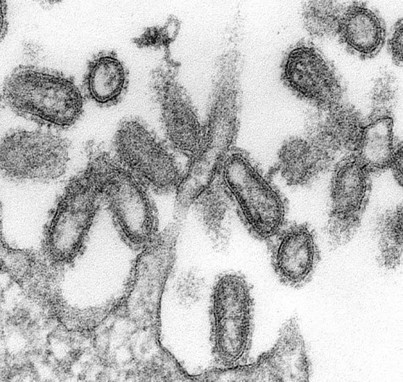
The 1918 influenza pandemic probably infected one-third of the world's population at the time — 500 million people. It killed between 50 million and 100 million; by contrast, Second World War deaths numbered around 60 million. Why is this catastrophe
Via Ed Rybicki
Robot scientist Eve can perform a wide range of biological assays, screening at a moderately high throughput rate (credit: Kevin Williams et al./J. R. Soc.
Via Gerd Moe-Behrens
From Canada, the ovillanta is a clever — and highly effective — mosquito trap made from the pests' favourite breeding spot.
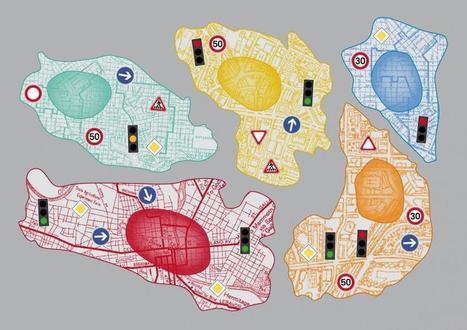
Cells regulate the uptake of nutrients and messenger cargos and their transport within the cell. This process is known as endocytosis and membrane traffic. Different cargos dock onto substrate specific receptors on the cell membrane. Special proteins such as kinases, GTPases and coats, activate specific entry routes and trigger the uptake of the receptors into the cell. For their uptake, the receptors and docked cargos become enclosed by the cell membrane. In the next steps, the membrane invaginates and becomes constricted. The resulting vesicle is guided via several distinct stations, cellular organelles, to its final destination in the cell. For her study, Dr. Prisca Liberali, senior scientist in the team of Professor Lucas Pelkmans, sequentially switched off 1200 human genes. Using automated high-throughput light microscopy and computer vision, she could monitor and compare 13 distinct transport paths involving distinct receptors and cellular organelles. Precise quantifications of thousands of single cells identified the genes required for the different transport routes. Surprisingly, sets of transport routes are co-regulated and coordinated in specific ways by different programs of regulatory control. Subsequently, Dr. Liberali calculated the hierarchical order within the genetic network and thereby identified the regulatory topology of cellular transport. "The transport into the cell and within the cells proceeds analogously to the cargo transport within a city" describes the scientist. "Like in a city, the traffic on the routes within a cell and their intersections is tightly regulated by traffic lights and signs to guide the cargo flow." Thanks to this unique quantitative map, the fine regulatory details of transport paths and processes within a cells could be mapped for the first time. Particularly the genes that encode for these traffic lights and switches are often de-regulated in disease. With this map, it is now possible to predict how this leads to traffic jams in the cells, causing the disease phenotype. Alternatively, since many drugs have been developed to target these traffic lights and switches, the map can be used to come up with possible drug combinations to target unwanted traffic, such as viruses, to the waste disposal system of the cell.
Via Dr. Stefan Gruenwald, burkesquires
A few years ago, I started looking online to fill in chapters of my family history that no one had ever spoken of.
A food poisoning bacterium may be implicated in MS, say US researchers.
IBM Wants to Predict Heart Disease Through Big Data Analytics
CloudTimes
Ultimately, health care industries hope to advance a smarter approach to care for patients with heart failure.
|
Zolgensma is a revolutionary gene therapy that can stop a deadly childhood condition called SMA in its tracks.It’s also one of the most expensive drugs in the world...
The pandemic has taught us that viruses are not easy to identify – and can spread like wildfire...
Results are in from the first phase 1 clinical trial for a Zika vaccine, and they are very promising. A new generation DNA-based Zika vaccine, developed by Wistar scientists in collaboration with Inovio Pharmaceuticals and GeneOne Life Science, was found to be safe and well tolerated by all study participants and was able to elicit an immune response against Zika, opening the door to further and larger trials to move this vaccine forward.
Via Integrated DNA Technologies
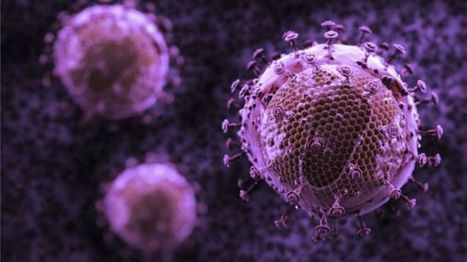
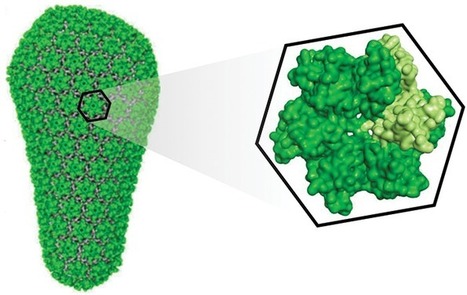
First potential new novel treatment for HIV in 10 years leverages the geometry of the HIV Capsid
The Indian vaccine, which protects against gastroenteritis caused by rotavirus, was tested in Niger.
Via Ed Rybicki
New study suggests HPV-related genital infection can cause cervical, anal, vulvar, and vaginal cancers.
We present a model of contagion that unifies and generalizes existing models of the spread of social influences and micro-organismal infections. Our model incorporates individual memory of exposure to a contagious entity (e.g., a rumor or disease), variable magnitudes of exposure (dose sizes), and heterogeneity in the susceptibility of individuals. Through analysis and simulation, we examine in detail the case where individuals may recover from an infection and then immediately become susceptible again (analogous to the so-called SIS model). We identify three basic classes of contagion models which we call \textit{epidemic threshold}, \textit{vanishing critical mass}, and \textit{critical mass} classes, where each class of models corresponds to different strategies for prevention or facilitation. We find that the conditions for a particular contagion model to belong to one of the these three classes depend only on memory length and the probabilities of being infected by one and two exposures respectively. These parameters are in principle measurable for real contagious influences or entities, thus yielding empirical implications for our model. We also study the case where individuals attain permanent immunity once recovered, finding that epidemics inevitably die out but may be surprisingly persistent when individuals possess memory. A generalized model of social and biological contagion
Peter Sheridan Dodds, Duncan J. Watts
Via Complexity Digest
Engineers from MIT have developed a substance that could be used as an artificial skin, long-lasting contact lens, drug-delivering bandage, and more.
Via Gerd Moe-Behrens
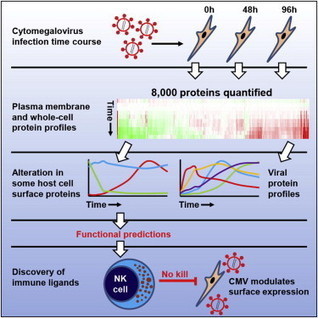
A systematic quantitative analysis of temporal changes in host and viral proteins throughout the course of a productive infection could provide dynamic insights into virus-host interaction. We developed a proteomic technique called “quantitative temporal viromics” (QTV), which employs multiplexed tandem-mass-tag-based mass spectrometry. Human cytomegalovirus (HCMV) is not only an important pathogen but a paradigm of viral immune evasion. QTV detailed how HCMV orchestrates the expression of >8,000 cellular proteins, including 1,200 cell-surface proteins to manipulate signaling pathways and counterintrinsic, innate, and adaptive immune defenses. QTV predicted natural killer and T cell ligands, as well as 29 viral proteins present at the cell surface, potential therapeutic targets. Temporal profiles of >80% of HCMV canonical genes and 14 noncanonical HCMV open reading frames were defined. QTV is a powerful method that can yield important insights into viral infection and is applicable to any virus with a robust in vitro model.
Via burkesquires
Life is a programming language, and molecular biologist Andrew Hessel thinks that it will be increasingly available to anyone interested in designing with the building blocks of life.
The NHS could be run more effectively if senior decision makers used simulation software to test the outcome of different approaches before rolling them out, according to a report out today.
Via Eugene Ch'ng
|
 Your new post is loading...
Your new post is loading...
 Your new post is loading...
Your new post is loading...
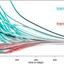























A very interesting ABM based approach for a stress-testing framework that could enable health authorities to rapidly identify bottlenecks in access to care and assess a healthcare networks resilience in the face of emergent situations. Well worth reading and investigating further.