Your new post is loading...
 Your new post is loading...
A new company offers a “gene vault” for doctors who want to add genomics to patient care. Genomic sequencing might be more common in medicine if doctors had a simple way to send for the test and keep track of the data. That’s the hope of Coriell Life Sciences in Camden, N.J., a startup that grew out of a partnership between the Coriell Institute for Medical Research and IBM. The company wants to facilitate the process of ordering, storing and interpreting whole-genome-sequence data for doctors. The company launched in January and is now working with different health-care providers to set up its service. “The intent is that the doctor would order a test like any other diagnostic test they order today,” says Scott Megill, president of Coriell Life Sciences. The company would facilitate sequencing the patient’s DNA (through existing sequencing companies such as Illumina or Ion Torrent), store it in its so-called gene vault, and act as the middleman between doctors and companies that offer interpretation services. Finally, “we will return the genetic result in the human readable form back to the electronic medical record so the doctor can read it and interpret it for the patient,” says Megill. “You need a robust software infrastructure for storing, analyzing, and presenting information,” says Jon Hirsch, who founded Syapse, a California-based company developing software to analyze biological data sets for diagnosing patients. “Until that gets built, you can generate all the data you want, but it’s not going to have any impact outside the few major centers of genomics medicine,” he says. The company will use a board of scientific advisors to guide them to the best interpretation programs available. “No one company is in position to interpret the entire genome for its meaning,” says Michael Christman, CEO of the Coriell Institute for Medical Research. “But by having one’s sequence in the gene vault, then the physician will be able to order interpretative engines, analogous to apps for the iPhone,” he says. Doctors could order an app to analyze a patient’s genome for DNA variants linked to poor drug response at one point, and later on, order another for variants linked to heart disease. The cloud-based workflow could help doctors in different locations take advantage of expert interpretations anywhere, says Christman. “This would allow a doctor who’s at a community clinic in Tulsa, Okla., order an interpretation of breast cancer sequences derived at Sloan Kettering,” he says.
Via Dr. Stefan Gruenwald
A new method of manufacturing short, single-stranded DNA molecules can solve many of the problems associated with current production methods. The new method can be of value to development of drugs consisting of DNA fragments and to DNA nanotechnology research. The novel technique for manufacturing short, single-stranded DNA molecules — or oligonucleotides — has been developed by researchers at Karolinska Institute in Sweden and Harvard University.
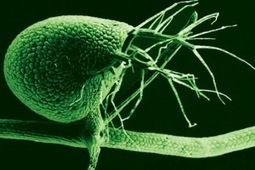
Only about two percent of the human genome contains genes. The other 98 percent has been likened to cosmology’s dark matter that fills the space between stars – there’s a lot of it, but nobody really knows what it does. Over the years scientists have put faith into the logic of evolution: if it’s there, it must serve a purpose. But a recent study shows that not all genomes are created equally. Unlike human genomes, the carnivorous bladderwort’s genome makes the most of its allotted bases having only an estimated 2 percent of non-coding DNA, or so-called “junk” DNA. The genome of the carnivorous bladderwort plant (Utricularia gibba) is minuscule compared to the human genome – 82,000 bases versus our near 3 billion. But while it’s small, the genome is extremely efficient. About 97 percent of its genome codes for an estimated 28,500 genes and the short sequences that control those genes. The authors of a study mapping the bladderwort genome surmise that, through many generations, the non-coding portion of the carnivorous bladderwort’s genome has been systematically removed, resulting in just 3 percent of non-coding DNA.
Healthy cells don't just happen. As they grow and divide, they need checks and balances to ensure they function properly while adapting to changing conditions around them. Researchers studying a set of proteins that regulate physiology, caloric restriction and aging have discovered another important role that one of them plays. SIRT4, one of seven sirtuin proteins, is known for controlling fuel usage from its post in the mitochondria, the cell's energy source. It responds to stressful changes in the availability of nutrients for the cell.
Using the technique that created Dolly the sheep, researchers from the RIKEN Center for Developmental Biology in Kobe, Japan, have identified a way to produce healthy mouse clones that live a normal lifespan and can be sequentially cloned indefinitely. In an experiment that started in 2005, the team led by Dr. Teruhiko Wakayama has used a technique called somatic cell nuclear transfer (SNCT) to produce 581 clones of one original "donor" mouse through 25 consecutive rounds of cloning.
Embracing the promise of personalized medicine, Fox Chase Cancer Center is offering a high-tech DNA test that can identify the genetic mutations driving an individual patient's cancer.
Via Brian Shields
What if you could improve your memory, become smarter and stronger, and live in an ageless disease-free body – just by taking a pill? Though this may sound like the stuff of science fiction, experts are developing a better understanding of our genetic mysteries, including the powerful influence that DNA wields on our lives. It's becoming clear that cancer, heart disease, diabetes, obesity; most mental disorders, and many other ailments, could all be the result of a clash between genes we inherited from our past, and today's modern environment.
I’m sick of reading about the dangers of the genome. There are lots of popular articles I could point to, but let’s start with a recent series in Time that included eight online features and the Dec. 13 cover story, ominously titled “The DNA Dilemma.” The series, written by Bonnie Rochman, is thoroughly reported, balanced, and full of fascinating personal stories about children whose genomes have been sequenced. It’s also timely: The primary question Rochman raises—how much information is too much information?—has been dominating commentaries about genetic testing in the medical literature. But this is the wrong question, or at least one that’s becoming increasingly irrelevant.
From cancer treatments to new devices to gene therapy, a look at six medical innovations that are poised to transform the way we fight disease.
Inducing cells to express telomerase, the enzyme which is supposed to slow down the metabolic clock, has enabled researchers boost the lifespan of mouse by 24% with a single treatment. The gene therapy acts on specific genes and is applied in adult life, not from the embryonic stage. Researchers at the Spanish National Cancer Research Center (CNIO) have demonstrated that the mouse lifespan can be extended by the application of one treatment acting directly on the animal’s genes in adult life. The therapy has been found to be safe and effective in mice. Adult and aged mice were treated with the gene therapy, delivering a rejuvenating effect. On average, the mice lived 24% longer. The older mice lived 13% longer. The therapy produces an appreciable improvement on the animal’s health and delayed the onset of age-related diseases. The genes were treated with a DNA-modified virus. The viral genes were replaced by those of the telomerase enzyme, which plays a key role in aging. Telomerase repairs the extreme ends of chromosomes, and slows the cells and therefore the body’s biological clock. There is a potential to develop a telomerase-based anti-aging gene therapy that won’t increase the risk of cancer. Telomeres are the caps that protect the end of chromosomes, but each time the cell divides, the telomeres get shorter until they lose all functionality. The cell then stops dividing or dies. Telomerase prevents telomeres from shortening or even rebuilding them. In most cells, telomerase is only active before birth. The cells of adult organisms contain no telomerase. There are some exceptions such as adult stem cells and cancer cells, which divide limitlessly and could be immortal. The risk of cancer tumor promotion is a risk that has set back telomerase-based anti-aging therapies. The kind of virus employed to carry the telomerase gene to the cells is very important. The viruses used is in this study have been successfully used in gene therapy treatment of hemophilia and eye disease. They are non-replicating viruses derived from others that are non-pathogenic in humans.
Via Dr. Stefan Gruenwald
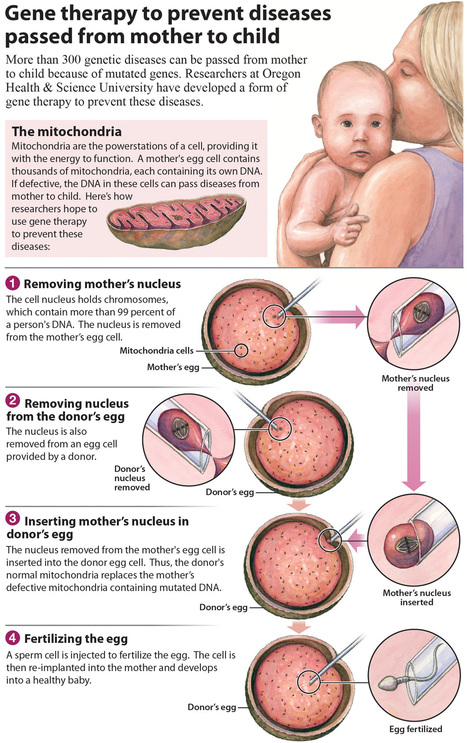
Scientists hope to prevent mitochondrial disease by removing chromosomes from the eggs of affected women, and putting them into donor eggs. Any children that would be born would not carry the mother’s mitochondrial mutations – but would have the mitochondrial DNA from the woman who donated her eggs.
UC Santa Barbara scientists Omar Saleh and Deborah Fygenson have created a dynamic gel made of DNA that mechanically responds to stimuli in much the same way that cells do. The project has potential applications in smart materials, artificial muscle, understanding cytoskeletal mechanics, research into nonequilibrium physics, and DNA nanotechnology.
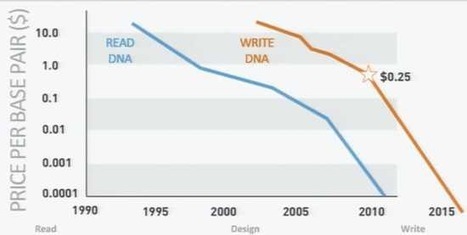
Today’s standard sequencing method uses an indirect approach to DNA sequencing based on fluorescently-tagged nucleotides, the building blocks of DNA. However, Ion Torrent has developed a simple, yet ingenious way to detect the sequence of nucleotides in a sample of DNA. Exploiting the fact that a hydrogen ion is released whenever a nucleotide attaches to a strand of DNA, sensors in Ion Torrent’s microchip have been designed to detect this chemical signature. Equality in gene sequencing ahead?
|
In a first of its kind ruling on human genes, a unanimous U.S. Supreme Court on Thursday decided that synthetically produced genetic material can be patented but naturally occurring DNA extracted from the human body cannot.
Scientists at The University of Nottingham have found that a genetic rogue element produced by sequences until recently considered “junk DNA” could promote cancer progression. The researchers, led by Dr Cristina Tufarelli, in the School of Graduate Entry Medicine and Health Sciences, discovered that the presence of this faulty genetic element — known as chimeric transcript LCT13 — is associated with the switching off of a known tumor suppressor gene (known as TFPI-2) whose expression is required to prevent cancer invasion and metastasis. Their findings suggest that LCT13 may be involved in switching off TFPI-2.
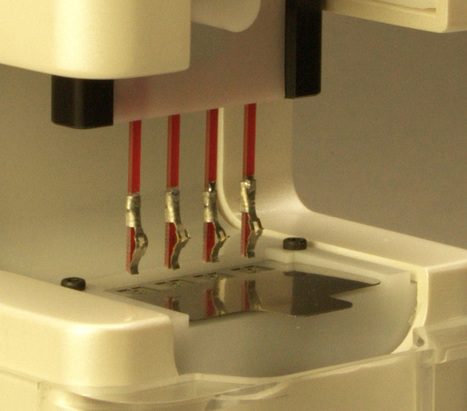
University of Washington engineers and NanoFacture, a Bellevue, Wash., company, have created a device that can extract human DNA from fluid samples in a simpler, more efficient and environmentally friendly way than conventional methods.
The device will give hospitals and research labs a much easier way to separate DNA from human fluid samples, which will help with genome sequencing, disease diagnosis and forensic investigations.
Stanford University bioengineers have taken computing beyond mechanics and electronics into the living realm of biology by creating the “transcriptor” — a biological transistor made from DNA and RNA.
In electronics, a transistor controls the flow of electrons along a circuit. Similarly, a transcriptor controls the flow of a specific protein, RNA polymerase, as it travels along a strand of DNA.
MIT researcher are developing a new dissolvable polymer film that replaces the vaccination needle with a pain-free patch.
According to a study carried out by a team led by María Blasco, the director of the Spanish National Cancer Research Centre (CNIO) and head of the Telomeres and Telomerase Group, a sustained lowering of food intake over time results in an increase of telomere length -- the ends of chromosomes -- in adult mice, which has a protective effect on the DNA and genetic material.
The notion that police can identify a suspect based on the tiniest drop of blood or trace of tissue has long been a staple of TV dramas, but scientists at Harvard have now taken the idea a step further. Using just a single human cell, they can reproduce an individual’s entire genome. The researchers developed a method — dubbed MALBAC, short for Multiple Annealing and Looping-based Amplification Cycles — that requires just one cell to reproduce an entire DNA molecule.
What worker bees do depends on how old they are. A worker a few days old will become a nurse bee that devotes herself to feeding larvae (brood), secreting beeswax to seal the cells that contain brood and attending to the queen.
Using a simple “drag-and-drop” computer interface and DNA self-assembly techniques, Parabon NanoLabs researchers have developed a new automated method of drug development that could reduce the time required to create and test medications, with the support of an NSF Technology Enhancement for Commercial Partnerships grant.
We may have sequenced the human and other species’ genomes, but we are still nowhere near predicting how this creates a living, breathing organism. Here’s why: In 2001, the Human Genome Project gave us an almost complete draft of the 3 billion letters in our DNA. We joined an elite club of species that have had their genome sequenced, one that is growing with every passing month. As our technologies and understanding advance, will we eventually be able to look at a pile of raw DNA sequence and glean all the workings of the organism it belongs to? Just as physicists can use the laws of mechanics to predict the motion of an object, can biologists use fundamental ideas in genetics and molecular biology to predict the traits and flaws of a body based solely on its genes? Could we pop a genome into a black box, and print out the image of a human? Or a fly? Or a mouse? Not easily. In complex organisms, some traits can be traced back to specific genes. If, for instance, you’re looking at a specific variant of the MC1R gene, chances are you’ve got a mammal in front of you, and it has red hair. Indeed, people have predicted that some Neanderthals were red-heads for precisely this reason. But beyond that, predicting if something is a mouse or a whale or a armadillo, we still can't do it. Bernhard Palsson from the University of California, San Diego agrees. “Sequencing a woolly mammoth will not predict its properties,” he says. “But you might be able to do a lot better with bacteria.” Their simpler and smaller genomes should in theory make it easier to predict the basic features of their metabolism, or whether they grow using oxygen or not. But even though we can sequence a bacterial genome in under a day, and for just $80, we would still struggle to determine important traits, like how good a disease-causing microbe is at infecting its host. Finding all the genes even in a small genome is hard. Earlier this year, scientists discovered a new gene in a flu virus whose genome consists of just 14,000 letters (small enough to fit into 100 tweets), and had been sequenced again and again. So it should be unsurprising that our own genome, with 3 billion letters, is full of errors and gaps, despite ostensibly being “complete”. In May, another group showed that the reference human genome is missing a gene that may have shaped the evolution of our large brains. “There’s no genome that is completely understood even in terms of the genes within it,” says Markus Covert from Stanford University. “Typically, no function is known for a fourth to a fifth of the genes, even in smaller genomes.”
Via Dr. Stefan Gruenwald
Imagine if DNA compilation was as easy to understand as Windows or iOS. Scientific study would no longer be necessary to engineer new combinations and just about anyone could drag and drop bits of genetic code into a workable sequence. Amirav-Drory wants to create a graphic user interface to empower people in just this way. His new software, Genome Compiler (free and available for download at www.genomecompiler.com), converts the various parts of a DNA sequence into easy-to-understand, and easily manipulable, icons.
|



 Your new post is loading...
Your new post is loading...