Your new post is loading...
 Your new post is loading...
Authors: Kun Dong, Fuqing Wu, Siqi Cheng, Shuai Li, Feng Zhang, Xinxin Xing, Xin Jin, Sheng Luo, Miao Feng, Rong Miao, Yanqi Chang, Shuang Zhang, Xiaoman You, Peiran Wang, Xin Zhang, Cailin Lei, Yulong Ren, Shanshan Zhu, Xiuping Guo, Chuanyin Wu, Dong-Lei Yang, Qibing Lin, Zhijun Cheng and Jianmin Wan.
Molecular Plant (2024)
Abstract: "Although both protein arginine methylation (PRMT) and jasmonate (JA) signaling are crucial for regulating plant development, the relationship between these processes in spikelet development control remains unclear. Here, we utilized CRISPR/Cas9 technology to generate two OsPRMT6a loss-of-function mutants exhibiting various abnormal spikelet structures. Additionally, we found that OsPRMT6a could methylate arginine residues in the JA signal repressors OsJAZ1 and OsJAZ7. Arginine methylation of OsJAZ1 increased the affinity of OsJAZ1 for the JA receptors OsCOI1a and OsCOI1b in the presence of jasmonates (JAs), subsequently promoting the ubiquitination of OsJAZ1 by the SCFOsCOI1a/OsCOI1b complex and degradation via the 26S proteasome. This process ultimately released OsMYC2, a core transcriptional regulator in the JA signaling pathway, to activate or repress JA-responsive genes, thereby maintaining normal plant (spikelet) development. However, in the osprmt6a-1 mutant, reduced arginine methylation of OsJAZ1 impaired the interaction between OsJAZ1 and OsCOI1a/OsCOI1b in the presence of JAs. As a result, OsJAZ1 proteins became more stable, repressing JA responses, thus causing the formation of abnormal spikelet structures. Moreover, we discovered that JA signaling reduced the OsPRMT6a mRNA level in an OsMYC2-dependent manner, thereby establishing a negative feedback loop to balance JA signaling. Furthermore, we found that OsPRMT6a-mediated arginine methylation of OsJAZ1 likely serves as a switch to tune JA signaling to maintain normal spikelet development under harsh environmental conditions such as high temperatures. Thus, our study established a direct molecular link between arginine methylation and the JA signaling pathway.
Abstract: "Cold stress severely restricts growth and development, reduces yields, and impairs quality in tomatoes (Solanum lycopersicum). Amylase-associated starch degradation and soluble sugar accumulation have been implicated in adaptation and resistance to abiotic stress. Here, we report a β-amylase (BAM) gene, SlBAM3, which plays a central role in tomato cold tolerance. The expression of SlBAM3 was triggered by cold stress. SlBAM3 knockout using the CRISPR/Cas9 system retarded starch degradation and reduced soluble sugar accumulation in tomato plants, eventually attenuating cold tolerance. Expression analysis revealed that the SlBAM3 transcript level was boosted by MeJA. Furthermore, MYC2, an essential component of the JA signaling pathway, could bind to the SlBAM3 promoter and directly activate SlBAM3 transcription, as revealed by yeast one-hybrid and dual LUC assays. In addition, the suppression of MYC2 resulted in increased starch accumulation, decreased soluble sugar content, and reduced tolerance to cold stress in tomato plants. Taken together, these findings demonstrate that JA positively regulates β-amylase-associated starch degradation through the MYC2-SlBAM3 module in tomato during cold stress. The results of the present work expand our understanding of the mechanisms underlying BAM gene activation and starch catabolism under cold stress. The regulatory module of SlBAM3 can be further utilized to breed tomato cultivars with enhanced cold tolerance."
Authors: Zachary Aldiss, Hannah Robinson, Yasmine Lam, Richard Dixon, Peter A. Crisp, Ian D. Godwin, Andrew Borrell, Lee Hickey and Karen Massel.
bioRxiv (2024)
Abstract: "Roots provide the critical interface where plants acquire nutrients and water, but our limited understanding of the genetic controls modulating root system architecture (RSA) in crop species constrains opportunities to develop future cultivars with improved root systems. However, there is vast knowledge of root developmental genes in model plant species, which has the potential to accelerate progress in crops with more complex genomes, particularly given that genome editing protocols are now available for most species. PIN-FORMED2 (PIN2) encodes a root specific polar auxin transporter, where its absence resulted in roots being unable to orient themselves using gravity, producing a significantly wider root system. To explore the role of PIN2 in a cereal crop, we used CRISPR/Cas9 editing to knockout of PIN2 in barley (Hordeum vulgare). Like Arabidopsis, the roots of barley pin2 loss-of-function mutants displayed an agravitropic response at seedling growth stages, resulting in a significantly shallower and wider root system at later growth stages. Notably, despite the significant change in RSA, there was no change in shoot architecture or total shoot biomass. We discuss the future challenges and opportunities to harness the PIN2 pathway to optimise RSA in crops for a range of production scenarios without a shoot trade-off."
Source: Agroscope Newsletter (15.02.2024)
Excerpts: "Agroscope has been granted approval by the Federal Office for the Environment for a field trial with spring barley. The focus is on a barley gene that has been disabled by new breeding techniques. The trial, which will be launched in spring 2024 on the Protected Site in Zurich-Reckenholz and will run for three years, aims to determine whether yields can be increased in this manner. The CKX2 gene is involved in the regulation of seed formation. Disabling this gene by means of a new breeding methods (CRISPR/Cas9 genome editing) brings about increased yields in rice and oilseed rape (see, below, ‘From rice to barley’).
"From rice to barley - Crop yield formation is complex and involves many different genes. However, Japanese researchers have discovered that the mutation of the CKX2 gene in rice has an unexpectedly significant effect on yield. Results were so convincing that they are now used in rice breeding. Research results show that genes corresponding to the CKX2 gene from rice also play a role e.g. in oilseed-rape yield formation. Therefore, it is reasonable to study this effect in further crops. In the best-case scenario, at the end of these trials on the Protected Site it will be possible to issue a recommendation as to whether breeders should disable one or both CKX2 genes in order to boost yields. What is certain, however, is that important information will be provided on the function of the CKX2 genes in barley – and hence further pieces of the puzzle will be available for a better understanding of yield formation."
Authors: XiangGuo Liu, Yang Liu, Ziqi Chen, Chuang Zhang, Jia Guo, Qing Liu, Yuejia Yin, Yang Hu, Hanchao Xia, Bingyang Li, Xiaopeng Sun and Yidan Li.
Research Square (2023)
Abstract: "Drought stress, a major plant abiotic stress, is capable of suppressing crop yield performance severely. However, the trade-off between crop drought tolerance and yield performance turns out to be significantly challenging in drought-resistant crop breeding. Several phytohormones (e.g., gibberellin (GA)) have been reported to play a certain role in plant drought response, which also take on critical significance in plant growth and development. In this study, the loss-of-function mutations of GA biosynthesis enzyme ZmGA20ox3 were produced using the CRISPR-Cas9 system in maize. As indicated by the result of two-year field trials, the above-mentioned mutants displayed semi-dwarfing phenotype with the decrease of GA1, and almost no yield loss was generated compared with wild-type (WT) plants. Interestingly, as revealed by the transcriptome analysis, differential expressed genes (DEGs) were notably enriched in abiotic stress progresses, and biochemical tests indicated the significantly increased ABA, JA, and DIMBOA levels in mutants, suggesting that ZmGA20ox3 may take on vital significance in stress response in maize. The in-depth analysis suggested that the loss function of ZmGA20ox3 can enhance drought tolerance in maize seedling, reduce Anthesis-Silking Interval (ASI) delay while decreasing the yield loss significantly in the field under drought conditions. The results of this study supported that regulating ZmGA20ox3 can improve plant height while enhancing drought resistance in maize, thus serving as a novel method for drought-resistant genetic improvement in maize."
Authors: Joy Y. Wang and Jennifer A. Doudna.
Science (2023)
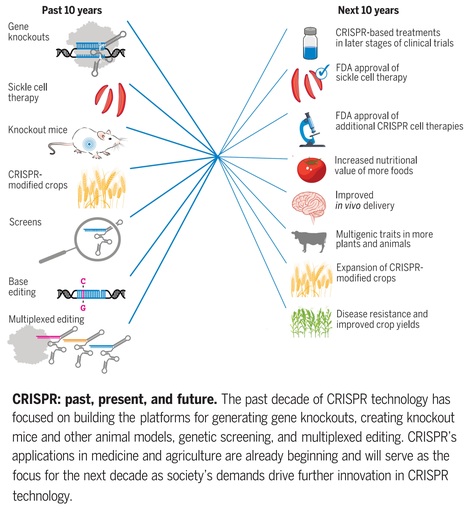
One-sentence summary: A review discusses the current state of CRISPR-mediated genetic manipulation in human cells, animals, and plants and considers its future potential.
Editor's view: A decade of CRISPR - In the decade since the publication of CRISPR-Cas9 as a genome-editing technology, the CRISPR toolbox and its applications have profoundly changed basic and applied biological research. Wang and Doudna now review the origins and utility of CRISPR-based genome editing, the successes and current limitations of the technology, and where innovation and engineering are needed. The authors describe important advances in the development of CRISPR genome-editing technology and make predictions about where the field is headed. They also highlight specific examples in medicine and agriculture that show how CRISPR is already affecting society, with exciting opportunities for the future. —DJ
Abstract: "The advent of clustered regularly interspaced short palindromic repeat (CRISPR) genome editing, coupled with advances in computing and imaging capabilities, has initiated a new era in which genetic diseases and individual disease susceptibilities are both predictable and actionable. Likewise, genes responsible for plant traits can be identified and altered quickly, transforming the pace of agricultural research and plant breeding. In this Review, we discuss the current state of CRISPR-mediated genetic manipulation in human cells, animals, and plants along with relevant successes and challenges and present a roadmap for the future of this technology."
Authors: Sayanti Mandal, Mimosa Ghorai, Uttpal Anand, Debleena Roy, Nishi Kant, Tulika Mishra, Abhijit Bhagwan Mane, Niraj Kumar Jha, Milan Kumar Lal, Rahul Kumar Tiwari, Manoj Kumar, Radha, Arabinda Ghosh, Rahul Bhattacharjee, Jarosław Proćków and Abhijit Dey.
Frontiers in Genetics (2022)
Abstract: "Over the last decade, remarkable progress has been made in our understanding the phytohormones, cytokinin’s (CKs) biosynthesis, perception, and signalling pathways. Additionally, it became apparent that interfering with any of these steps has a significant effect on all stages of plant growth and development. As a result of their complex regulatory and cross-talk interactions with other hormones and signalling networks, they influence and control a wide range of biological activities, from cellular to organismal levels. In agriculture, CKs are extensively used for yield improvement and management because of their wide-ranging effects on plant growth, development and physiology. One of the primary targets in this regard is cytokinin oxidase/dehydrogenase (CKO/CKX), which is encoded by CKX gene, which catalyses the irreversible degradation of cytokinin. The previous studies on various agronomically important crops indicated that plant breeders have targeted CKX directly. In recent years, prokaryotic clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein 9 (Cas9) system has been increasingly used in editing the CKO/CKX gene and phenomenal results have been achieved. This review provides an updated information on the applications of CRISPR-based gene-editing tools in manipulating cytokinin metabolism at the genetic level for yield improvement. Furthermore, we summarized the current developments of RNP-mediated DNA/transgene-free genomic editing of plants which would broaden the application of this technology. The current review will advance our understanding of cytokinins and their role in sustainably increase crop production through CRISPR/Cas genome editing tool."
|
Authors: Shammi Akter, Oscar Castaneda-Méndez and Jesús Beltrán
Current Opinion in Biotechnology (2024)
Highlights • Several challenges remain for the widespread use of synthetic biology in crops. • Synthetic gene expression circuitry is advancing in model plants. • Rapid enzyme discovery and testing are needed for plant metabolic engineering.
Abstract: "Plant synthetic biology (Plant SynBio) is an emerging field with the potential to enhance agriculture, human health, and sustainability. Integrating genetic tools and engineering principles, Plant SynBio aims to manipulate cellular functions and construct novel biochemical pathways to develop plants with new phenotypic traits, enhanced yield, and be able to produce natural products and pharmaceuticals. This review compiles research efforts in reprogramming plant developmental and biochemical pathways. We highlight studies leveraging new gene expression toolkits to alter plant architecture for improved performance in model and crop systems and to produce useful metabolites in plant tissues. Furthermore, we provide insights into the challenges and opportunities associated with the adoption of Plant SynBio in addressing complex issues impacting agriculture and human health."
Authors: Connor Tansley, Nicola J. Patron, and Sarah Guiziou
ACS Synthetic Biology (2024)
Abstract: "Many plant species are grown to enable access to specific organs or tissues, such as seeds, fruits, or stems. In some cases, a value is associated with a molecule that accumulates in a single type of cell. Domestication and subsequent breeding have often increased the yields of these target products by increasing the size, number, and quality of harvested organs and tissues but also via changes to overall plant growth architecture to suit large-scale cultivation. Many of the mutations that underlie these changes have been identified in key regulators of cellular identity and function. As key determinants of yield, these regulators are key targets for synthetic biology approaches to engineer new forms and functions. However, our understanding of many plant developmental programs and cell-type specific functions is still incomplete. In this Perspective, we discuss how advances in cellular genomics together with synthetic biology tools such as biosensors and DNA-recording devices are advancing our understanding of cell-specific programs and cell fates. We then discuss advances and emerging opportunities for cell-type-specific engineering to optimize plant morphology, responses to the environment, and the production of valuable compounds."
"Regulation der Korngröße bei Weizen entdeckt"
Quelle: Pflanzenforschung.de (12.03.2024)
Durch Ausschalten des TabHLH489-Gens mit der Genschere CRISPR/Cas erhöht sich die Körnergröße und Ertrag von Brotweizen.
Auszüge: "Weizen ist die weltweit bedeutendste Getreideart nach Mais und die Erntemenge beträgt ca. 800 Millionen Tonnen im laufenden Wirtschaftsjahr 2023/24. Um die Versorgungssicherheit für die weiterwachsende Weltbevölkerung zu gewährleisten, sind in den kommenden Jahren deutliche Ertragssteigerungen notwendig. Chinesische Forscher haben dazu einen neuen Ansatz gefunden. Durch Genom-Editierung mit der Genschere CRISPR/Cas konnte das Team Weizenvarianten erzeugen, die höhere Erträge liefern. Im Fachmagazin Plant Biotechnology Journal erläutern die Forscher:innen, dass das Gen TabHLH489 eine Schlüsselrolle für die Korngröße spielt."
"Das Ausschalten des Gens für TabHLH489 und seinen homologen Genen mittels der Genschere CRISPR/Cas führte zu einer Zunahme der Kornlänge und des Korngewichts, während die Überexpression zu einer Abnahme beider Werte führte. Darüber hinaus konnten sie einen ganzen Regulationskomplex aufdecken: TaSnRK1α1, die α-katalytische Untereinheit des pflanzlichen Energiesensors SnRK1, interagierte mit TabHLH489 und phosphorylierte den Transkriptionsfaktor, um seinen Abbau zu induzieren und so die Entwicklung des Weizenkorns zu fördern. Eine Zuckerbehandlung induzierte ebenfalls die Anreicherung des TaSnRK1α1-Proteins und führte so zu einer Senkung des TabHLH489-Proteinspiegels..... Auch Brassinosteroide steuern die Korngröße - Darüber hinaus konnten die Forscher:innen zeigen, dass das Pflanzenhormon Brassinosteroid (BR) die Kornentwicklung fördert, indem es die TabHLH489-Expression über den Transkriptionsfaktor BRASSINAZOLE RESISTANT1 (BZR1) verringert. Eine weitere wichtige Beobachtung war, dass natürliche Variationen in der Promotorregion von TabHLH489 die BZR1-Bindungsfähigkeit beeinträchtigen und dadurch die TabHLH489-Expression beeinflussen. Insgesamt konnte die Studie damit den grundlegenden Regulationsmechanismus für die Korngröße bei Weizen aufklären: das TaSnRK1α1-TabHLH489-Regulationsmodul, das auch BR- und Zuckersignale integriert. Damit steht nun der Weg offen, gezielt Weizensorten mit erhöhter Korngröße und Ertrag zu züchten."
Authors: Sunny Ahmar and Damian Gruszka.
Trends in Biochemical Sciences (2023)
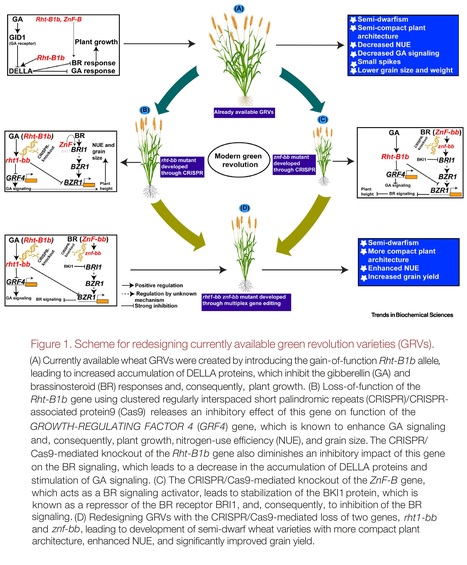
Abstract: "A modern green revolution is needed to ensure global food security. Recently, Song et al. reported a new strategy to create high-yielding, semi-dwarf wheat varieties with improved nitrogen-use efficiency by inhibiting brassinosteroid (BR) signaling through clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein9 (Cas9)-mediated knockout of the ZnF-B gene encoding a zinc-finger RING-type E3 ligase."
In: Phys.org
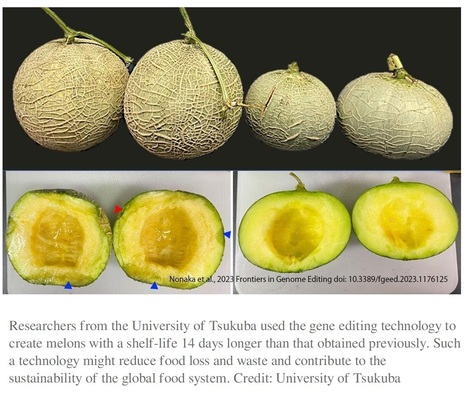
Excerpts: "The gaseous plant hormone ethylene has been long known to promote fruit ripening and plays a certain role in shelf-life. In a study published in Frontiers in Genome Editing, researchers performed gene editing using the Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/Cas9 system via modification of the ethylene synthesis pathway in the Japanese luxury melon (Cucumis melo var. reticulatus "Harukei-3") to increase its shelf life."
"In this study, CmACO1 was selected as a target of gene editing and attempted to introduce mutations in the gene. The harvested melons exhibited no foreign genes and the mutations induced were inherited for at least two generations. In the non-gene-edited line (wild type), ethylene generation was observed in the fruit 14 days post-harvest, the rind turned yellow, and the flesh softened. However, in the genome-edited mutant, ethylene generation was reduced to one-tenth of that in the wild type, with the skin color remaining green and the fruit remaining firm. This indicates that introducing CmACO1 mutation via gene editing enhanced the shelf life of the melons."
Authors: Huaxia Shi, Ying Xu, Na Tian, Ming Yang and Fu-Sen Liang.
Nature Communications (2022)
Editor's view: RNA modifications, including N6-methyladenosine (m6A), have been reported to regulate fundamental RNA processes and properties, and directly linked to various human diseases. Here, the authors develop a chemically inducible and reversible RNA m6A modification editing platform integrating chemically induced proximity (CIP) and CRISPR methods.
Abstract: "RNA modifications, including N6-methyladenosine (m6A), have been reported to regulate fundamental RNA processes and properties, and directly linked to various human diseases. Methods enabling temporal and transcript/locus-specific editing of specific RNA modifications are essential, but still limited, to dissect the dynamic and context-dependent functions of these epigenetic modifications. Here, we develop a chemically inducible and reversible RNA m6A modification editing platform integrating chemically induced proximity (CIP) and CRISPR methods. We show that m6A editing can be temporally controlled at specific sites of individual RNA transcripts by the addition or removal of the CIP inducer, abscisic acid (ABA), in the system. By incorporating a photo-caged ABA, a light-controlled version of m6A editing platform can be developed. We expect that this platform and strategy can be generally applied to edit other RNA modifications in addition to m6A."
|



 Your new post is loading...
Your new post is loading...