Your new post is loading...
 Your new post is loading...
Authors: Jorge Hernández-García, Antonio Serrano-Mislata, María Lozano-Quiles, Cristina Úrbez, María A. Nohales, Noel Blanco-Touriñán, Huadong Peng, Rodrigo Ledesma-Amaro and Miguel A. Blázquez.
PNAS (2024)
Significance: DELLA proteins are plant-specific transcriptional hubs integrating environmental signals with endogenous cues. In order to regulate downstream processes, DELLAs modulate the activity of hundreds of transcription factors (TFs) and transcriptional regulators in various ways. Here, we describe the molecular mechanism underlying DELLA coactivator function. We show that DELLAs act as transcriptional activators by interacting with the Mediator complex subunit MED15. This interaction is necessary to regulate a specific subset of DELLA-dependent responses that are mediated by transcriptional coactivation, but not those regulated by TF sequestration. We further show that this mechanism is present in bryophyte DELLAs and thus represents a conserved mechanism of DELLA function in land plants.
Abstract: "DELLA proteins are negative regulators of the gibberellin response pathway in angiosperms, acting as central hubs that interact with hundreds of transcription factors (TFs) and regulators to modulate their activities. While the mechanism of TF sequestration by DELLAs to prevent DNA binding to downstream targets has been extensively documented, the mechanism that allows them to act as coactivators remains to be understood. Here, we demonstrate that DELLAs directly recruit the Mediator complex to specific loci in Arabidopsis, facilitating transcription. This recruitment involves DELLA amino-terminal domain and the conserved MED15 KIX domain. Accordingly, partial loss of MED15 function mainly disrupted processes known to rely on DELLA coactivation capacity, including cytokinin-dependent regulation of meristem function and skotomorphogenic response, gibberellin metabolism feedback, and flavonol production. We have also found that the single DELLA protein in the liverwort Marchantia polymorpha is capable of recruiting MpMED15 subunits, contributing to transcriptional coactivation. The conservation of Mediator-dependent transcriptional coactivation by DELLA between Arabidopsis and Marchantia implies that this mechanism is intrinsic to the emergence of DELLA in the last common ancestor of land plants."
Authors: Ting Li, Yongqin Wang, Annelore Natran, Yi Zhang, Hao Wang, Kangxi Du, Peng Qin, Hua Yuan, Weilan Chen, Bin Tu, Dirk Inzé and Marieke Dubois.
New Phytologist (2024)
Abstract: "Gibberellic acid (GA) plays a central role in many plant developmental processes and is crucial for crop improvement. DELLA proteins, the core suppressors in the GA signaling pathway, are degraded by GA via the 26S proteasomal pathway to release the GA response. However, little is known about the phosphorylation-mediated regulation of DELLA proteins. In this study, we combined GA response assays with protein–protein interaction analysis to infer the connection between Arabidopsis thaliana DELLAs and the C-TERMINAL DOMAIN PHOSPHATASE-LIKE 3 (CPL3), a phosphatase involved in the dephosphorylation of RNA polymerase II. We show that CPL3 directly interacts with DELLA proteins and promotes DELLA protein stability by inhibiting its degradation by the 26S proteasome. Consequently, CPL3 negatively modulates multiple GA-mediated processes of plant development, including hypocotyl elongation, flowering time, and anthocyanin accumulation. Taken together, our findings demonstrate that CPL3 serves as a novel regulator that could improve DELLA stability and thereby participate in GA signaling transduction."
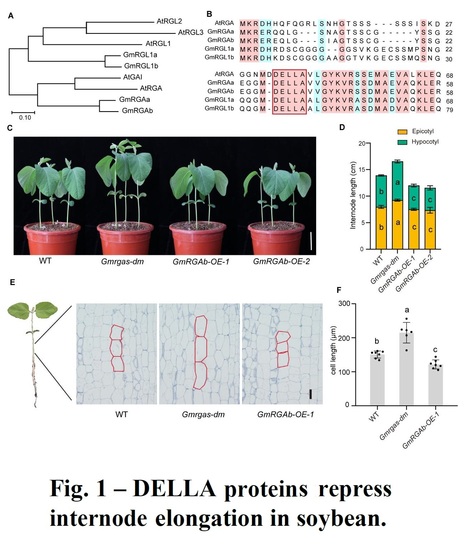
Authors: Zhuang Li, Qichao Tu, Xiangguang Lyu, Qican Cheng, Ronghuan Ji, Chao Qin, Jun Liu, Bin Liu, Hongyu Li and Tao Zhao.
The Crop Journal (2024)
Abstract: "Plant height influences plant architecture, lodging resistance, and yield performance. It is modulated by gibberellic acid (GA) metabolism and signaling. DELLA proteins, acting as central repressors of GA signaling, integrate various environmental and hormonal signals to regulate plant growth and development in Arabidopsis. We examined the role of two DELLA proteins, GmRGAa and GmRGAb, in soybean plant height control. Knockout of these proteins led to longer internodes and increased plant height, primarily by increasing cell elongation. GmRGAs functioned under different light conditions, including red, blue, and far-red light, to repress plant height. Interaction studies revealed that GmRGAs interacted with the blue light receptor GmCRY1b. Consistent with this, GmCRY1b partially regulated plant height via GmRGAs. Additionally, DELLA proteins were found to stabilize the protein GmSTF1/2, a key positive regulator of photomorphogenesis. This stabilization led to increased transcription of GmGA2ox-7b and subsequent reduction in plant height. This study enhances our understanding of DELLA-mediated plant height control, offering Gmrgaab mutants for soybean structure and yield optimization.
Authors: Jingyun Lu, Guifang Zhang, Chao Ma, Yao Li, Chuyan Jiang, Yaru Wang, Bingjie Zhang, Rui Wang, Yuexuan Qiu, Yanxing Ma, Yangchao Jia, Cai-Zhong Jiang, Xiaoming Sun, Nan Ma, Yunhe Jiang and Junping Gao.
The Plant Cell (2024)
One-sentence summary: An ethylene-induced F-box protein, RhSAF, accelerates petal senescence by destabilizing the gibberellic acid receptor RhGID1.
Abstract: "Roses are among the most popular ornamental plants cultivated worldwide for their great economic, symbolic, and cultural importance. Nevertheless, rapid petal senescence markedly reduces rose (Rosa hybrida) flower quality and value. Petal senescence is a developmental process tightly regulated by various phytohormones. Ethylene accelerates petal senescence, while gibberellic acid (GA) delays this process. However, the molecular mechanisms underlying the crosstalk between these phytohormones in the regulation of petal senescence remain largely unclear. Here, we identified SENESCENCE-ASSOCIATED F-BOX (RhSAF), an ethylene-induced F-box protein gene encoding a recognition subunit of the SCF-type E3 ligase. We demonstrated that RhSAF promotes degradation of the GA receptor GIBBERELLIN INSENSITIVE DWARF1 (RhGID1) to accelerate petal senescence. Silencing RhSAF expression delays petal senescence, while suppressing RhGID1 expression accelerates petal senescence. RhSAF physically interacts with RhGID1s and targets them for ubiquitin/26S proteasome-mediated degradation. Accordingly, ethylene-induced RhGID1C degradation and RhDELLA3 accumulation are compromised in RhSAF-RNAi lines. Our results demonstrate that ethylene antagonizes GA activity through RhGID1 degradation mediated by the E3 ligase RhSAF. These findings enhance our understanding of the phytohormone crosstalk regulating petal senescence and provide insights for improving flower longevity."
Authors: Baoshan Xian, Muhammad Saad Rehmani, Yueni Fan, Xiaofeng Luo, Ranran Zhang, Jiahui Xu, Shaowei Wei, Lei Wang, Juan He, Aigen Fu and Kai Shu.
New Phytologist (2024)
Abstract: "Abscisic acid (ABA) and gibberellins (GA) antagonistically mediate several biological processes, including seed germination, but the molecular mechanisms underlying ABA/GA antagonism need further investigation, particularly any role mediated by a transcription factors module. Here, we report that the DELLA protein RGL2, a repressor of GA signaling, specifically interacts with ABI4, an ABA signaling enhancer, to act as a transcription factor complex to mediate ABA/GA antagonism. The rgl2, abi3, abi4 and abi5 mutants rescue the non-germination phenotype of the ga1-t. Further, we demonstrate that RGL2 specifically interacts with ABI4 to form a heterodimer. RGL2 and ABI4 stabilize one another, and GA increases the ABI4-RGL2 module turnover, whereas ABA decreases it. At the transcriptional level, ABI4 enhances the RGL2 expression by directly binding to its promoter via the CCAC cis-element, and RGL2 significantly upregulates the transcriptional activation ability of ABI4 toward its target genes, including ABI5 and RGL2. Abscisic acid promotes whereas GA inhibits the ability of ABI4-RGL2 module to activate transcription, and ultimately ABA and GA antagonize each other. Genetic analysis demonstrated that both ABI4 and RGL2 are essential for the activity of this transcription factor module. These results suggest that the ABI4-RGL2 module mediates ABA/GA antagonism by functioning as a double agent."
Authors: Zhuang Li, Xiangguang Lyu, Hongyu Li, Qichao Tu, Tao Zhao, Jun Liu and Bin Liu.
Nature Communications (2024)
Editor's view: This study provides insights into how shade induces leaf senescence in soybean. The reduction of blue light intensity deactivates GmCRY1s, leading to the degradation of GmRGAs and the upregulation of WRKY100, ultimately promoting leaf senescence.
Abstract: "Leaf senescence is a crucial trait that has a significant impact on crop quality and yield. Previous studies have demonstrated that light is a key factor in modulating the senescence process. However, the precise mechanism by which plants sense light and control senescence remains largely unknown, particularly in crop species. In this study, we reveal that the reduction in blue light under shading conditions can efficiently induce leaf senescence in soybean. The blue light receptors GmCRY1s rather than GmCRY2s, primarily regulate leaf senescence in response to blue light signals. Our results show that GmCRY1s interact with DELLA proteins under light-activated conditions, stabilizing them and consequently suppressing the transcription of GmWRKY100 to delay senescence. Conversely, LBL reduces the interaction between GmCRY1s and the DELLA proteins, leading to their degradation and premature senescence of leaves. Our findings suggest a GmCRY1s-GmDELLAs-GmWRKY100 regulatory cascade that is involved in mediating LBL-induced leaf senescence in soybean, providing insight into the mechanism of how light signals regulate leaf senescence. Additionally, we generate GmWRKY100 knockout soybeans that show delayed leaf senescence and improved yield under natural field conditions, indicating potential applications in enhancing soybean production by manipulating the leaf senescence trait."
Authors: Zizhao Xie, Liang Jin, Ying Sun, Chenghang Zhan, Siqi Tang, Tian Qin, Nian Liu and Junli Huang.
Plant Communications (2024)
Abstract: "The crosstalk between gibberellin (GA) and abscisic acid (ABA) signaling is crucial for balancing plant growth and adaption to environmental stress. Nevertheless, the molecular mechanism of mutual antagonism still needs to be well understood. In this study, we find that knockout of rice NAC (NAM, ATAF1/2, CUC2) transcription factor gene OsNAC120 inhibits plant growth while enhances drought tolerance, but its overexpression lines act in an opposite way. Exogenous GA can rescue the semi-dwarf phenotype of osnac120 mutants, and further study shows that OsNAC120 promotes GA biosynthesis through transcriptionally activating GA biosynthetic genes OsGA20ox1 and OsGA20ox3. DELLA protein SLENDER RICE1 (SLR1) interacts with OsNAC120 and impedes its transactivation ability, while GA treatment can remove the inhibition of transactivation activity caused by SLR1. On the other hand, OsNAC120 negatively regulates rice drought tolerance by repressing ABA-induced stomatal closure. Mechanistic investigation illustrates that OsNAC120 inhibits ABA biosynthesis via transcriptional repression of ABA biosynthetic genes OsNCED3 and OsNCED4. Rice OSMOTIC STRESS/ABA-ACTIVATED PROTEIN KINASE 9 (OsSAPK9) physically interacts with OsNAC120 and mediates its phosphorylation, which results in OsNAC120 degradation. ABA treatment accelerates OsNAC120 degradation and reduces its transactivation activity. Together, our findings provide the evidence that OsNAC120 plays critical roles in balancing GA-mediated growth and ABA-induced drought tolerance in rice. The research will help us understand the mechanisms underlying the “trade-off” between plant growth and stress tolerance, and engineer stress-resistant and high-yielding crops."
Authors: Jorge Hernández-García, Antonio Serrano-Mislata, María Lozano-Quiles, Cristina Úrbez, María A. Nohales, Noel Blanco-Touriñán, Huadong Peng, Rodrigo Ledesma-Amaro and Miguel A. Blázquez.
bioRxiv (2023)
Abstract: "DELLA proteins are negative regulators of the gibberellin response pathway in angiosperms, acting as central hubs that interact with hundreds of transcription factors and regulators to modulate their activities. While the mechanism of transcription factor sequestration by DELLAs to prevent DNA binding to downstream targets has been extensively documented, the mechanism that allows them to act as co-activators remains to be understood. Here, we demonstrate that DELLAs directly recruit the Mediator complex to specific loci in Arabidopsis, facilitating transcription. This recruitment involves DELLA amino-terminal domain and the conserved MED15 KIX domain. Accordingly, partial loss of MED15 function mainly disrupted processes known to rely on DELLA co-activation capacity; including cytokinin-dependent regulation of meristem function and skotomorphogenic response, gibberellin metabolism feedback, and flavonol production. We have also found that the single DELLA protein in the liverwort Marchantia polymorpha is capable of recruiting MpMED15 subunits, contributing to transcriptional co-activation. The conservation of Mediator-dependent transcriptional co-activation by DELLA between Arabidopsis and Marchantia implies that this mechanism is intrinsic to the emergence of DELLA in the last common ancestor of land plants."
Authors: Junjie Li, Qi Li, Wentao Wang, Xinran Zhang, Chen Chu, Xintian Tang, Bo Zhu, Lizhong Xiong, Yu Zhao and Dao-Xiu Zhou.
The EMBO Journal (2023)
Synopsis: DELLA proteins are master repressors of gibberellin (GA) signaling, but the molecular basis of this repression is not well understood. This work shows that one DELLA forms a tripartite complex with Polycomb-repressive complex 2 (PRC2) and histone deacetylase HDA702 to repress GA-inducible genes by establishing a silent chromatin state in rice. The rice DELLA protein SLENDER RICE1 (SLR1) interacts with PRC2 and HDA702. SLR1 is required for H3K27 trimethylation (H3K27me3) at a subset of PRC2 targets. GA signaling increases H3K9 acetylation (H3K9ac) and decreases H3K27me3. GA signaling dissociates PRC2 and HDA702 from target genes.
Abstract: "DELLA proteins are master regulators of gibberellic acid (GA) signaling through their effects on gene expression. Enhanced DELLA accumulation in rice and wheat varieties has greatly contributed to grain yield increases during the green revolution. However, the molecular basis of DELLA-mediated gene repression remains elusive. In this work, we show that the rice DELLA protein SLENDER RICE1 (SLR1) forms a tripartite complex with Polycomb-repressive complex 2 (PRC2) and the histone deacetylase HDA702 to repress downstream genes by establishing a silent chromatin state. The slr1 mutation and GA signaling resulted in dissociation of PRC2 and HDA702 from GA-inducible genes. Loss-of-function or downregulation of the chromatin regulators impaired SLR1-dependent histone modification and gene repression. Time-resolved analysis of GA signaling revealed that GA-induced transcriptional activation was associated with a rapid increase of H3K9ac followed by H3K27me3 removal. Collectively, these results establish a general epigenetic mechanism for DELLA-mediated gene repression and reveal details of the chromatin dynamics during transcriptional activation stimulated by GA signaling."
Authors: Imani Madison, Lydia Gillan, Jasmine Peace, Flavio Gabrieli, Lisa Van den Broeck, Jacob L Jones and Rosangela Sozzani.
Journal of Experimental Botany (2023)
Abstract: "Phosphorus is essential to plant growth and agricultural crop yields, yet the challenges associated with phosphorus fertilization in agriculture, such as aquatic runoff pollution and poor phosphorus bioavailability, are increasingly difficult to manage. Comprehensively understanding the dynamics of phosphorus uptake and signaling mechanisms will inform the development of strategies to address these issues. This review describes regulatory mechanisms used by specific tissues in the root apical meristem to sense and uptake phosphate from the rhizosphere. The major regulatory mechanisms and related hormone crosstalk underpinning phosphate starvation responses, cellular phosphate homeostasis, and plant adaptations to phosphate starvation are also discussed in this review. In addition, this review overviews the major mechanism of plant systemic phosphate starvation responses. Finally, this review discusses recent promising genetic engineering strategies for improving crop phosphorus use and computational approaches that may help further design strategies for improved plant phosphate acquisition. The mechanisms and approaches presented in this review include a wide variety of species not only including Arabidopsis thaliana but also including crop species such as Oryza sativa (rice), Glycine max (soybean), and Triticum aestivum (wheat) to address both general and species-specific mechanisms and strategies. The aspects of phosphorus deficiency responses and recently employed strategies of improving phosphate acquisition that are detailed in this review may provide insights on the mechanisms or phenotypes that may be targeted in efforts to improve crop phosphorus content and plant growth in low phosphorus soils."
Authors: Xu Huang, Hao Tian, Jeongmoo Park, Dong-Ha Oh, Jianhong Hu, Rodolfo Zentella, Hong Qiao, Maheshi Dassanayake and Tai-Ping Sun.
Nature Plants (2023)
Editor's view: Genetic and ChIP–seq analyses of missense DELLA mutants reveal a role of the DELLA PFYRE subdomain in H2A binding, which stabilizes the transcription factor–DELLA–H2A complexes at the target chromatin for global transcription reprogramming.
Abstract: "The DELLA genes, also known as ‘Green Revolution’ genes, encode conserved master growth regulators that control plant development in response to internal and environmental cues. Functioning as nuclear-localized transcription regulators, DELLAs modulate expression of target genes via direct protein–protein interaction of their carboxy-terminal GRAS domain with hundreds of transcription factors (TFs) and epigenetic regulators. However, the molecular mechanism of DELLA-mediated transcription reprogramming remains unclear. Here by characterizing new missense alleles of an Arabidopsis DELLA, repressor of ga1-3 (RGA), and co-immunoprecipitation assays, we show that RGA binds histone H2A via the PFYRE subdomain within its GRAS domain to form a TF–RGA–H2A complex at the target chromatin. Chromatin immunoprecipitation followed by sequencing analysis further shows that this activity is essential for RGA association with its target chromatin globally. Our results indicate that, although DELLAs are recruited to target promoters by binding to TFs via the LHR1 subdomain, DELLA–H2A interaction via the PFYRE subdomain is necessary to stabilize the TF–DELLA–H2A complex at the target chromatin. This study provides insights into the two distinct key modular functions in DELLA for its genome-wide transcription regulation in plants."
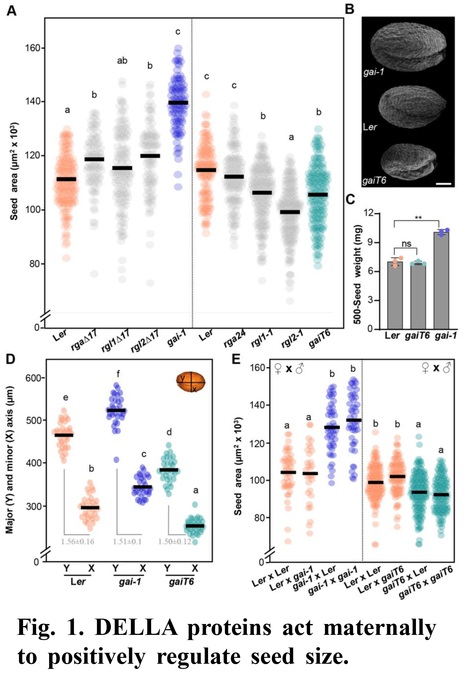
Authors: Maria Dolores Gomez, Isabel Cored, Daniela Barro-Trastoy, Joaquin Sanchez-Matilla, Pablo Tornero and Miguel A. Perez-Amador.
Development (2023)
Abstract: "Human and animal nutrition is mainly based on seeds. Seed size is a key factor affecting seed yield and has thus been one of the primary objectives of plant breeders since the domestication of crop plants. Seed size is coordinately regulated by signals of maternal and zygotic tissues that control the growth of the seed coat, endosperm, and embryo. Here, we provide novel evidence of the role of DELLA proteins, key repressors of gibberellin responses, in the maternal control of seed size. The gain-of-function della mutant gai-1 produces larger seeds due to an increase in the cell number in ovule integuments. This leads to an increase in ovule size and, in turn, an increase in seed size. Moreover, DELLA activity promotes increased seed size by inducing the transcriptional activation of AINTEGUMENTA, a genetic factor that controls cell proliferation and organ growth, in the ovule integuments of gai-1. Overall, our results point to DELLA proteins as new players in control of seed size and suggest that modulation of the DELLA-dependent pathway could be used to improve crop yield."
Authors: Chunyu Zhang, Mingyang Jian, Weijun Li, Xiani Yao, Cuirong Tan, Qian Qian, Yilong Hu, Liu Xu and Xingliang Hou.
The Plant Cell (2023)
Abstract: "Gibberellin (GA) plays a key role in floral induction by activating the expression of floral integrator genes in plants, but the epigenetic regulatory mechanisms underlying this process remain unclear. Here, we show that BRAHMA (BRM), a core subunit of the chromatin remodeling SWI/SNF complex that functions in various biological processes by regulating gene expression, is involved in GA signaling-mediated flowering via the formation of the DELLA-BRM-NF-YC module in Arabidopsis (Arabidopsis thaliana). DELLA, BRM, and NF-YC transcription factors interact with each other, and DELLA proteins promote the physical interaction between BRM and NF-YC proteins. This impairs the binding of NF-YCs to SOC1, a major floral integrator gene, to inhibit flowering. On the other hand, DELLA proteins also facilitate the binding of BRM to SUPPRESSOR OF OVEREXPRESSION OF CONSTANS1 (SOC1). The GA-induced degradation of DELLA proteins disturbs the DELLA-BRM-NF-YC module, prevents BRM from inhibiting NF-YCs, and decreases the DNA binding ability of BRM, which promotes the deposition of H3K4me3 on SOC1 chromatin, leading to early flowering. Collectively, our findings show that BRM is a key epigenetic partner of DELLA proteins during the floral transition. Moreover, they provide molecular insights into how GA signaling coordinates an epigenetic factor with a transcription factor to regulate the expression of a flowering gene and flowering in plants."
|
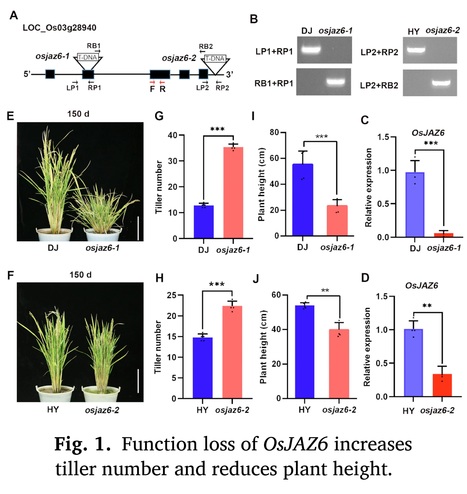
Authors: Wanmin Wang, Zizhao Xie, Yuanyuan Wu, Ying Sun, Chenghang Zhan, Liang Jin and Junli Huang.
Environmental and Experimental Botany (2024)
Highlights: • OsJAZ6 modulates rice tillering and drought response by integrating JA with GA signaling. • OsJAZ6 controls the tiller bud growth but not formation. • OsJAZ6 interacts with SLR1 to promote its degradation, which further destabilizes MOC1. • OsJAZ6 and SLR1 have opposite functions in regulating rice tiller bud growth and drought tolerance.
Abstract: "Jasmonic acid (JA) plays crucial functions during plant growth and stress response, but its roles and regulatory mechanism in plant branching remain largely unknown. Rice basal branching (tillering) is an essential agronomic trait that affects crop production. Here, we report that OsJAZ6, the repressor of JA signaling, negatively modulates rice tillering and drought stress tolerance. Loss-of-function mutants of OsJAZ6 exhibit a significant increase in tiller number, while OsJAZ6ΔJas-overexpression lines produce fewer tillers than wild-type plants. Further investigations show that function loss of OsJAZ6 promotes the tiller bud growth rather than formation. Mechanistic studies show that OsJAZ6 interacts with rice DELLA/SLR1 (SLENDER RICE 1), a transcription repressor of gibberellin (GA) signaling, and the interaction promotes SLR1 degradation, which further facilitates the degradation of rice tillering regulator MOC1 (MONOCULM 1), thereby inhibiting the tiller bud growth. In agreement, the slr1 mutant exhibits fewer tillers than wild type. Consistently, application of JA promotes the growth of tiller bud and thus increases the tiller number, while GA treatment results in opposite result. Meanwhile, osjaz6 mutants display enhanced drought tolerance, coupled with increased JA sensitivity, while the slr1 mutant shows the reverse behavior. Collectively, our data demonstrate that OsJAZ6 negatively modulates rice tillering as well as drought stress tolerance by destabilizing SLR1 protein. Our data shed light on the regulatory mechanism of controlling the tiller development and drought stress response in rice by the JA-OsJAZ6-SLR1 module."
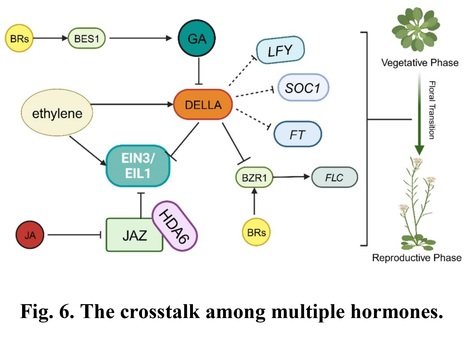
Authors: Xiaoxiao Li, Chuyu Lin, Chenghao Lan and Zeng Tao.
Journal of Experimental Botany (2024)
Abstract: "The timing of the developmental transition from the vegetative to the reproductive stages is critical for angiosperm and fine-tuned by the integration of endogenous factors and external environmental cues to ensure proper and successful reproduction. Plants have evolved sophisticated mechanisms to response to diverse environmental or stress signals, which may be mediated by plant hormones which coordinate their flowering time. Endogenous and exogenous phytohormones such as gibberellin (GA), auxin, cytokinin (CK), jasmonate (JA), abscisic acid (ABA), ethylene (ET), brassinosteroids (BR) and the cross-talk among them are critical for the precise regulating of flowering time. Recent studies on the model flowering plant Arabidopsis thaliana revealed that diverse transcription factors and epigenetic regulators play key roles in the phytohormones that regulate floral transition. This review aims to summarize current knowledge on the genetic and epigenetic mechanisms that underlying the phytohormone control of floral transition in Arabidopsis, offering insights into how these processes are regulated and their implications for plant biology."
Authors: Peng He, Liping Zhu, Xin Zhou, Xuan Fu, Yu Zhang, Peng Zhao, Bin Jiang, Huiqin Wang and Guanghui Xiao.
Developmental Cell (2024)
Editor's view: He et al. identified two signaling cascades, GhSLR1-GhZFP8-GhSDCP1- GhPIF3 and GhSLR1-GhBLH1-GhKCS12, which translate the gibberellins signal to regulate cotton fiber cell elongation. This finding can be potentially used in the field to improve cotton fiber quality.
Highlight: • Gibberellins promotes fiber cell elongation by degrading GhSLR1 • GhSLR1 interacts with two transcription factors, GhZFP8 and GhBLH1 • GhBLH1 activates the transcription of GhKCS12 and enhances VLCFA biosynthesis • GhZFP8 activates the expression of GhPIF3 by upregulating GhSDCP1 in cotton
Abstract: "The agricultural green revolution spectacularly enhanced crop yield through modification of gibberellin (GA) signaling. However, in cotton, the GA signaling cascades remain elusive, limiting our potential to cultivate new cotton varieties and improve yield and quality. Here, we identified that GA prominently stimulated fiber elongation through the degradation of DELLA protein GhSLR1, thereby disabling GhSLR1’s physical interaction with two transcription factors, GhZFP8 and GhBLH1. Subsequently, the resultant free GhBLH1 binds to GhKCS12 promoter and activates its expression to enhance VLCFAs biosynthesis. With a similar mechanism, the free GhZFP8 binds to GhSDCP1 promoter and activates its expression. As a result, GhSDCP1 upregulates the expression of GhPIF3 gene associated with plant cell elongation. Ultimately, the two parallel signaling cascades synergistically promote cotton fiber elongation. Our findings outline the mechanistic framework that translates the GA signal into fiber cell elongation, thereby offering a roadmap to improve cotton fiber quality and yield."
Authors: Guy Golan, Jacob Weiner, Yusheng Zhao and Thorsten Schnurbusch.
New Phytologist (2024)
Abstract: "How plants distribute biomass among organs influences resource acquisition, reproduction and plant–plant interactions, and is essential in understanding plant ecology, evolution, and yield production in agriculture. However, the genetic mechanisms regulating allocation responses to the environment are largely unknown. We studied recombinant lines of wheat (Triticum spp.) grown as single plants under sunlight and simulated canopy shade to investigate genotype-by-environment interactions in biomass allocation to the leaves, stems, spikes, and grains. Size-corrected mass fractions and allometric slopes were employed to dissect allocation responses to light limitation and plant size. Size adjustments revealed light-responsive alleles associated with adaptation to the crop environment. Combined with an allometric approach, we demonstrated that polymorphism in the DELLA protein is associated with the response to shade and size. While a gibberellin-sensitive allelic effect on stem allocation was amplified when plants were shaded, size-dependent effects of this allele drive allocation to reproduction, suggesting that the ontogenetic trajectory of the plant affects the consequences of shade responses for allocation. Our approach provides a basis for exploring the genetic determinants underlying investment strategies in the face of different resource constraints and will be useful in predicting social behaviours of individuals in a crop community.
Authors: Eilon Shani, Peter Hedden and Tai-ping Sun.
Plant Physiology (2024)
One-sentence summary: A historical overview of gibberellin research showcases important advances in our understanding of gibberellin metabolism, perception, signaling, and transport.
Abstract: "It has been almost a century since biologically active gibberellin (GA) was isolated. Here, we give a historical overview of the early efforts in establishing the GA biosynthesis and catabolism pathway, characterizing the enzymes for GA metabolism, and elucidating their corresponding genes. We then highlight more recent studies that have identified the GA receptors and early GA signaling components (DELLA repressors and F-box activators), determined the molecular mechanism of DELLA-mediated transcription reprogramming, and revealed how DELLAs integrate multiple signaling pathways to regulate plant vegetative and reproductive development in response to internal and external cues. Finally, we discuss the GA transporters and their roles in GA-mediated plant development."
Authors: Zhen Liu, Xiao-Feng Cheng, Ying-Ning Zou, Anoop Kumar Srivastava, Mashael Daghash Alqahtani and Qiang-Sheng Wu.
Environmental and Experimental Botany (2024)
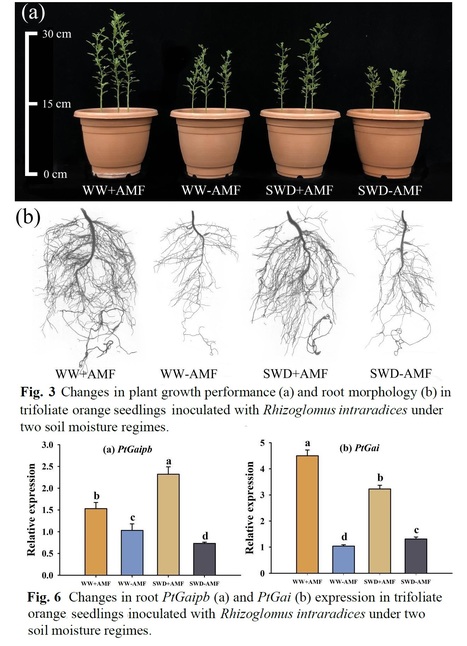
Highlights • AMF boosted plant growth as well as POD and CAT activity in roots under drought. • Nine GA species were detected in roots, with GA4 only found in AMF roots. • AMF reduced GA3, but increased GA5, GA9, and GA15 levels under drought. • AMF up-regulated the expression of PtCPS, PtPKS, PtKO, PtGA20ox, PtGA3ox, and PtGA2ox under drought. • AMF increased the expression of the DELLA genes PtGaipb and PtGai under drought.
Abstract: "Gibberellins (GAs), an important endogenous hormone, serve a crucial regulatory role in plants’ resistance to environmental stress, whereas it is unclear whether and how arbuscular mycorrhizal fungi (AMF) regulate the profile of GAs in plants exposed to soil water deficit (SWD). This study aimed to analyze how Rhizoglomus intraradices inoculation affected plant growth performance, antioxidant enzyme activities, and the composition, synthesis, metabolism, and signaling pathways of GAs in the roots of trifoliate orange (Poncirus trifoliata) under a 10-week SWD. Although SWD decreased both root AMF colonization and soil hyphal length, mycorrhizal plants nevertheless outperformed non-mycorrhizal plants in terms of growth performance, as well as peroxidase (70.0%) and catalase (149.2%) activities, indicating greater drought tolerance. A total of nine GAs compositions were detected in the roots, with bioactive GA4 exclusively found in AMF roots. AMF colonization significantly reduced bioactive GA3 levels by 37.9%, while it increased inactive GA8, GA9, and GA15 levels under SWD by 522.2%, 100.0%, and 2500.0%. Inoculation with AMF also up-regulated the expression of PtCPS, PtPKS, and PtKO in GA synthesis pathways of roots and the expression of PtGA20ox, PtGA3ox, and PtGA2ox in GA metabolisms under SWD. In addition, the expression of DELLA genes PtGaipb and PtGai associated with GA signaling pathways was further up-regulated under SWD by AMF inoculation. It is concluded that AMF regulated the composition, synthesis, and deactivation of GAs in roots to promote plant growth, along with an increase in PtGaipb and PtGai expression to possibly initiate mycorrhizal signaling pathways in response to SWD."
Authors: Catarina Neves, Beatriz Ribeiro, Rute Amaro, Jesús Expósito, Jérôme Grimplet and Ana Margarida Fortes.
Horticulture Research (2023)
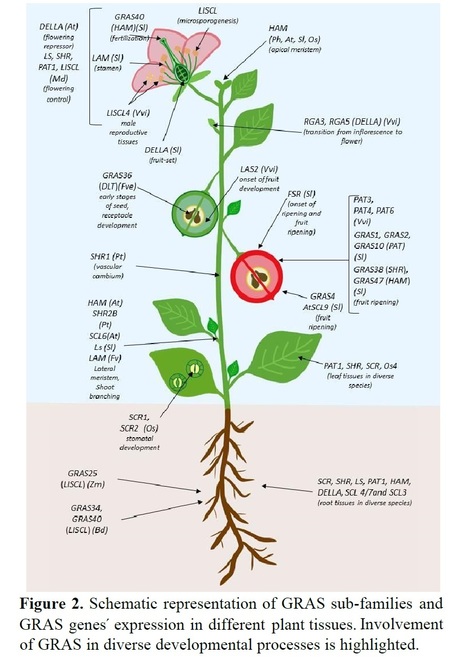
Abstract: "The plant-specific family of GRAS transcription factors has been wide implicated in the regulation of transcriptional reprogramming associated with a diversity of biological functions ranging from plant development processes to stress responses. Functional analyses of GRAS transcription factors supported by in silico structural and comparative analyses are emerging and clarifying the regulatory networks associated with their biological roles. In this review, a detailed analysis of GRAS proteins´ structure and biochemical features as revealed by recent discoveries indicated how these characteristics may impact subcellular location, molecular mechanisms, and function. Nomenclature issues associated with GRAS classification into different subfamilies in diverse plant species even in the presence of robust genomic resources are discussed, in particular how it affects assumptions of biological function. Insights into the mechanisms driving evolution of this gene family and how genetic and epigenetic regulation of GRAS contributes to subfunctionalization are provided. Finally, this review debates challenges and future perspectives on the application of this complex but promising gene family for crop improvement to cope with challenges of environmental transition."
Authors: Hao Wu, Qi He, Bing He, Shuyi He, Longjun Zeng, Longbo Yang, Hong Zhang, Zhaoran Wei, Xingming Hu, Jiang Hu, Yong Zhang, Lianguang Shang, Suikang Wang, Peng Cui, Guosheng Xiong, Qian Qian and Quan Wang.
The Plant Cell (2023)
Abstract: "The elimination of seed shattering was a key step in rice (Oryza sativa) domestication. In this paper, we show that increasing the gibberellic acid (GA) content or response in the abscission region enhanced seed shattering in rice. We demonstrate that SLENDER RICE1 (SLR1), the key repressor of GA signaling, could physically interact with the rice seed shattering-related transcription factors QTL of seed shattering on chromosome 1 (qSH1), Oryza sativa HOMEOBOX 15 (OSH15), and SUPERNUMERARY BRACT GENE (SNB). Importantly, these physical interactions interfered with the direct binding of these three regulators to the lignin biosynthesis gene 4-COUMARATE:COENZYME A LIGASE 3 (4CL3), thereby de-repressing its expression. De-repression of 4CL3 led to increased lignin deposition in the abscission region, causing reduced rice seed shattering. Importantly, we also show that modulating GA content could alter the degree of seed shattering to increase harvest efficiency. Our results reveal that the “Green Revolution” phytohormone GA is important for regulating rice seed shattering, and we provide an applicable breeding strategy for high-efficiency rice harvesting."
Authors: Matías Ezequiel Pereyra, Cecilia Costigliolo Rojas, Anne F. Jarrell, Austin S. Hovland, Stephen A. Snipes, Punita Nagpal, David Alabadí, Miguel A. Blázquez, Rodrigo A. Gutiérrez, Jason W. Reed, William M. Gray and Jorge José Casal.
PNAS (2023)
Significance: Nitrate increases the yield potential of crops but, by enhancing plant stature, it also exacerbates the risk of lodging. During the Green Revolution, the introduction of dwarfing genes alleviated this problem, but at the expense of nitrogen use efficiency. By using the hypocotyl of Arabidopsis thaliana as a model, we elucidate the mechanisms by which nitrate promotes plant stature. Hypocotyl elongation responds to nitrate upshifts rather than to the actual levels of the nutrient. Nitrate sensing increases the nuclear abundance of selected transcription factors, which increase the expression of numerous members of the SAUR gene family involved in the promotion of cell expansion. The pathway reveals potential targets to genetically reduce plant stature without the drawbacks of current approaches.
Abstract: "Nitrate supply is fundamental to support shoot growth and crop performance, but the associated increase in stem height exacerbates the risks of lodging and yield losses. Despite their significance for agriculture, the mechanisms involved in the promotion of stem growth by nitrate remain poorly understood. Here, we show that the elongation of the hypocotyl of Arabidopsis thaliana, used as a model, responds rapidly and persistently to upshifts in nitrate concentration, rather than to the nitrate level itself. The response occurred even in shoots dissected from their roots and required NITRATE TRANSPORTER 1.1 (NRT1.1) in the phosphorylated state (but not NRT1.1 nitrate transport capacity) and NIN-LIKE PROTEIN 7 (NLP7). Nitrate increased PHYTOCHROME INTERACTING FACTOR 4 (PIF4) nuclear abundance by posttranscriptional mechanisms that depended on NRT1.1 and phytochrome B. In response to nitrate, PIF4 enhanced the expression of numerous SMALL AUXIN-UP RNA (SAUR) genes in the hypocotyl. The growth response to nitrate required PIF4, positive and negative regulators of its activity, including AUXIN RESPONSE FACTORs, and SAURs. PIF4 integrates cues from the soil (nitrate) and aerial (shade) environments adjusting plant stature to facilitate access to light."
Authors: Dengan Xu, Yingjie Bian, Xumei Luo, Chenfei Jia, Qianlin Hao, Xiuling Tian, Qiang Cao, Wei Chen, Wujun Ma, Zhongfu Ni, Xiangdong Fu, Zhonghu He, Xianchun Xia and Shuanghe Cao.
Development (2023)
Summary: Transgenic, micromorphological, dynamic and multi-omic analyses in wheat reveal the underlying mechanism of the Green Revolution gene Rht-B1b in modulation of plant architecture and yield component traits.
Abstract: "The utilization of reduced plant height genes Rht-B1b and Rht-D1b, encoding homeologous DELLA proteins, led to the wheat Green Revolution (GR). However, the specific functions of GR genes in yield determination and the underlying regulatory mechanisms remained unknown. Here, we validated that Rht-B1b, as a representative of GR genes, affects plant architecture and yield component traits. Upregulation of Rht-B1b reduced plant height, leaf size and grain weight, but increased tiller number, tiller angle, spike number per unit area, and grain number per spike. Dynamic investigations showed that Rht-B1b increased spike number by improving tillering initiation rather than outgrowth, and enhanced grain number by promoting floret fertility. Rht-B1b reduced plant height by reducing cell size in the internodes, and reduced grain size or weight by decreasing cell number in the pericarp. Transcriptome analyses uncovered that Rht-B1b regulates many homologs of previously reported key genes for given traits and several putative integrators for different traits. These findings specify the pleiotropic functions of Rht-B1b in improving yield and provide new insights into the regulatory mechanisms underlying plant morphogenesis and yield formation."
Author: Lucas Frungillo.
The Plant Cell (2023)
Excerpts: "The brown planthopper (BPH), Nilaparvata lugens, is a major threat to rice (Oryza sativa)."
"Because the phytohormone gibberellic acid (GA) regulates plant growth and development, the authors sought to investigate the impact of BPH feeding on GA levels in rice. Hormonal profiling by liquid chromatography–mass spectrometry (LC-MS) revealed lower levels of GA bioactive forms in BPH-attacked rice compared to control plants. Accordingly, overexpression of the GA catabolic genes GA2ox3 and GA2ox7 resulted in lower levels of bioactive GA and reduced growth as compared to wild-type plants (see Figure), suggesting that herbivore attack triggers the catabolism of bioactive GAs in rice."
"Collectively, Jin et al. (2023) provide compelling evidence that the MYC2-GA2ox module orchestrates JA-GA hormonal crosstalk in rice during defense responses against BPH attack. The mechanistic insights presented in this study open exciting new directions for the investigation of hormonal crosstalk in plants and provide a framework for improving resistance to pests without yield penalties in rice.
Authors: Peipei Xu, Jinbo Hu, Haiying Chen and Weiming Cai.
Cell Reports (2023)
Editor's view: Xu et al. show that KAR signaling protein SMAX1 inhibits seed germination under weak light conditions by regulating gibberellin (GA) biosynthesis. SMAX1 inhibits GA3ox2 gene expression and DELLA proteins interact with SMAX1 to regulate its transcriptional activity, indicating a crosstalk between KAR and GA signaling in regulating Arabidopsis seed germination.
Highlights • KAR signaling mutants have lower germination percentage under weak light • SMAX1 functions as an auto-regulated transcription factor • The interactions of SMAX1 with DELLA proteins regulate SMAX1 transcriptional activity • SMAX1 can inhibit the GA3ox2 gene expression, which is a key enzyme in GA biosynthesis
Abstract: "Karrikins (KARs) were first identified as a class of small-molecule chemicals derived from smoke that promote seed germination. However, the implied mechanism is still not well understood. Here, we find that KAR signaling mutants have a lower germination percentage than that of wild type under weak light conditions, and KARs promote seed germination through transcriptional activation of gibberellin (GA) biosynthesis via SMAX1. SMAX1 interacts with the DELLA proteins REPRESSOR of ga1-3-LIKE 1 (RGL1) and RGL3. The interaction enhances the transcriptional activity of SMAX1 and inhibits GIBBERELLIN 3-oxidase 2 (GA3ox2) gene expression. The KAR signaling mutant seed germination defect under weak light is partially rescued by exogenous application of GA3 or by GA3ox2 overexpression, and the rgl1 rgl3 smax1 triple mutant exhibits higher germination rates under weak light than the smax1 mutant. Thus, we show a crosstalk between KAR and GA signaling pathways via a SMAX1-DELLA module in regulating seed germination in Arabidopsis."
|



 Your new post is loading...
Your new post is loading...