Your new post is loading...
 Your new post is loading...
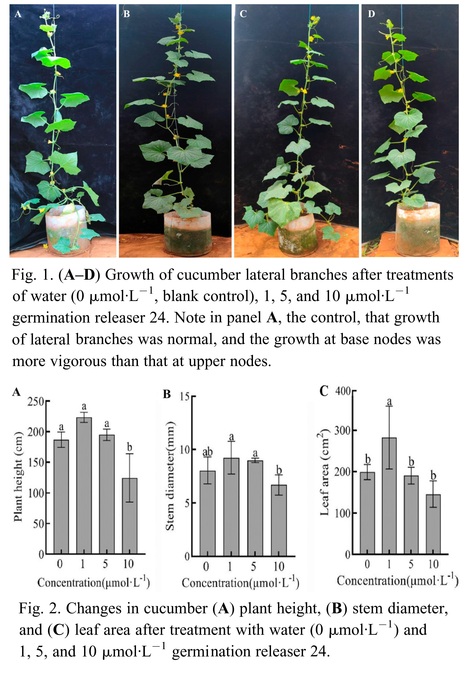
Authors: Tian Su, Ziwei Li, Yinghua Zhang, Junqiang Xu and Bin Xu.
Journal of the American Society for Horticultural Science (2024)
Abstract: "Cucumber (Cucumis sativus L.) belongs to the cucumber genus of the Cucurbitaceae family, and the selection of cultivars with minimal or no lateral branches can enhance the cultivation management efficiency. The growth of lateral branches is inhibited by strigolactone. To investigate the regulatory mechanism of strigolactone on the lateral branch development in cucumber, the cultivar LZ1 exhibiting multiple lateral branches was selected as the experimental material. The axillae of the plants were infiltrated with 1, 5, and 10 μmol·L−1 germination releaser 24 (GR24) at the four- to five-leaf stage. It was identified that 1 μmol·L−1 GR24 exhibited the most potent inhibitory effect on cucumber lateral branches. Additionally, exogenous strigolactone decreased the auxin content in the apical bud and axillae and increased the auxin content in the stem. This inhibited polar auxin transport in the axillary bud and promoted polar auxin transport in the apical bud. The content of strigolactone in the axilla region of cucumbers was elevated, whereas the synthesis and expression of cytokinin in the same area were reduced. A low concentration of GR24 induced the expression of cucumber branched 1 (csbrc1), whereas a high concentration of GR24 downregulated the expression of cucumber lateral suppressor (cscls) and blind (csblind), which inhibited the growth of cucumber lateral branches."
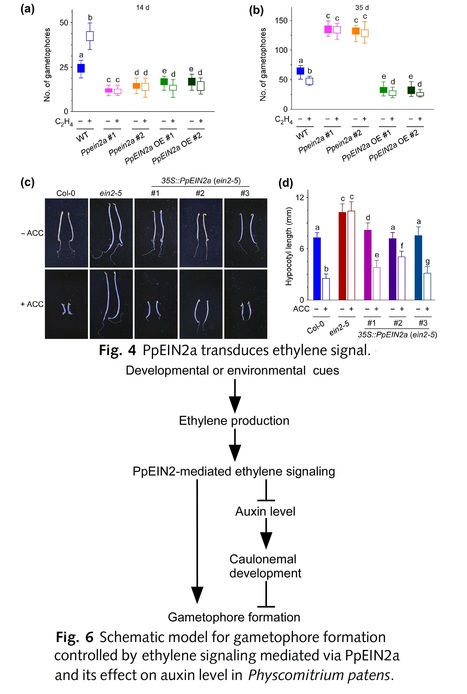
Authors: Yidong Wang, Lanlan Jiang, Dongdong Kong, Jie Meng, Meifang Song, Wenxiu Cui, Yaqi Song, Xiaofan Wang, Jiao Liu, Rui Wang, Yikun He, Caren Chang and Chuanli Ju.
New Phytologist (2024)
Abstract: "The conquest of land by plants was concomitant with, and possibly enabled by, the evolution of three-dimensional (3D) growth. The moss Physcomitrium patens provides a model system for elucidating molecular mechanisms in the initiation of 3D growth. Here, we investigate whether the phytohormone ethylene, which is believed to have been a signal before land plant emergence, plays a role in 3D growth regulation in P. patens. We report ethylene controls 3D gametophore formation, based on results from exogenously applied ethylene and genetic manipulation of PpEIN2, which is a central component in the ethylene signaling pathway. Overexpression (OE) of PpEIN2 activates ethylene responses and leads to earlier formation of gametophores with fewer gametophores produced thereafter, phenocopying ethylene-treated wild-type. Conversely, Ppein2 knockout mutants, which are ethylene insensitive, show initially delayed gametophore formation with more gametophores produced later. Furthermore, pharmacological and biochemical analyses reveal auxin levels are decreased in the OE lines but increased in the knockout mutants. Our results suggest that evolutionarily, ethylene and auxin molecular networks were recruited to build the plant body plan in ancestral land plants. This might have played a role in enabling ancient plants to acclimate to the continental surfaces of the planet."
Authors: Chang Su, Andrzej Kokosza, Xiaonan Xie, Aleš Pěnčík, Youjun Zhang, Pasi Raumonen, Xueping Shi, Sampo Muranen, Melis Kucukoglu Topcu, Juha Immanen, Risto Hagqvist, Omid Safronov, Juan Alonso-Serra, Gugan Eswaran, Mirko Pavicic Venegas, Karin Ljung, Sally Ward, Ari Pekka Mähönen, Kristiina Himanen, Jarkko Salojärvi, Alisdair R. Fernie, Ondřej Novák, Ottoline Leyser, Wojtek Pałubicki, Ykä Helariutta and Kaisa Nieminen.
PNAS (2023)
Significance What makes a tree a tree instead of a bush? Through a candidate gene approach, we identified a natural bush-like (short and highly branching) SL-deficient birch mutant, kanttarelli, with an early STOP codon in an essential SL biosynthesis gene, BpMAX1. The number of higher-order branches was increased in the mutant and in phenocopying transgenic RNAi -lines. Intriguingly, the auxin concentration formed a gradient along the main stem in the WT, with more auxin in the uppermost internodes and less toward the base, whereas in the transgenic line this gradient was absent. Mathematical modeling showed that this difference in auxin distribution may result from the differing architectures. Our results could be applied in the breeding of trees with an optimized architecture.
Abstract: "Due to their long lifespan, trees and bushes develop higher order of branches in a perennial manner. In contrast to a tall tree, with a clearly defined main stem and branching order, a bush is shorter and has a less apparent main stem and branching pattern. To address the developmental basis of these two forms, we studied several naturally occurring architectural variants in silver birch (Betula pendula). Using a candidate gene approach, we identified a bushy kanttarelli variant with a loss-of-function mutation in the BpMAX1 gene required for strigolactone (SL) biosynthesis. While kanttarelli is shorter than the wild type (WT), it has the same number of primary branches, whereas the number of secondary branches is increased, contributing to its bush-like phenotype. To confirm that the identified mutation was responsible for the phenotype, we phenocopied kanttarelli in transgenic BpMAX1::RNAi birch lines. SL profiling confirmed that both kanttarelli and the transgenic lines produced very limited amounts of SL. Interestingly, the auxin (IAA) distribution along the main stem differed between WT and BpMAX1::RNAi. In the WT, the auxin concentration formed a gradient, being higher in the uppermost internodes and decreasing toward the basal part of the stem, whereas in the transgenic line, this gradient was not observed. Through modeling, we showed that the different IAA distribution patterns may result from the difference in the number of higher-order branches and plant height. Future studies will determine whether the IAA gradient itself regulates aspects of plant architecture."
Authors: Rameshwar Sharma and Yellamaraju Sreelakshmi.
Journal of Experimental Botany (2023)
Abstract: "Axillary buds (ABs) are dormant buds located in the leaf axils of plants, which have the potential to develop into branches or flowers under appropriate conditions. At the molecular-genetic level, the miR156/SPL/SPB module regulates the development of ABs in plants, thus influencing plant architecture. Auxins are plant hormones that regulate various aspects of plant growth and development, including AB activity. Barrera-Rojas et al. (2023) show that suppressing AB outgrowth elevates auxin levels and lowers GOBLET expression, probably by suppressing its transcription by SBP15. Their findings provide insights into the regulation of AB outgrowth and tomato shoot architecture."
Authors: Jun Wang, Jing Huang, Jinlin Bao, Xizhi Li, Liang Zhu and Jian Jin.
Molecular Plant (2023)
Abstract: "Plant architecture and panicle architecture are two critical agronomic traits that largely affect the yield of rice (Oryza sativa L.). PROSTRATE GROWTH 1 (PROG1) encodes a key C2H2-type zinc-finger transcription factor and has pleiotropic effects on the regulation of both plant and panicle architecture, thereby influencing the grain yield. However, the molecular mechanisms through which PROG1 controls plant and panicle architecture remain unclear. In this study, we have demonstrated the direct binding of PROG1 to the LAZY 1 (LA1) promoter and its role as a repressor of LA1. Conversely, LA1 acts as a repressor of PROG1 by directly binding to the PROG1 promoter. These two genes play antagonistic roles in shaping plant architecture by regulating both tiller angle and tiller number. Furthermore, our findings reveal that PROG1 controls panicle architecture through direct binding to the intragenic regulatory regions of OsGIGANTEA (OsGI) and subsequent activation of its expression. Overall, our study has identified two crucial targets of PROG1, LA1 and OsGI, shedding light on the primary molecular mechanisms underlying plant and panicle architecture control by PROG1. This provides valuable insights into the regulation of key domestication-related traits in rice and identifies potential targets for future high-yield rice breeding."
Authors: Craig L. Cowling, Linkan Dash and Dior R. Kelley.
Journal of Experimental Botany (2023)
Abstract: "Phytohormones play a central role in plant development and environmental responses. Auxin is a classical hormone that is required for organ formation, tissue patterning, and defense responses. Auxin pathways have been extensively studied across numerous land plant lineages, including bryophytes and eudicots. In contrast, our understanding of the roles of auxin in maize morphogenesis and immune responses are limited. Here, we will review evidence for auxin-mediated processes in maize and describe promising areas for future research in the auxin field. Several recent transcriptomic and genetic studies have demonstrated that auxin is a key influencer of both vegetative and reproductive development in maize (namely roots, leaves and kernels). Auxin signaling has been implicated in both maize shoot architecture and immune responses through genetic and molecular analyses of the conserved co-repressor RAMOSA ENHANCER LOCUS2. Polar auxin transport is linked to maize drought responses, root growth, shoot formation, and leaf morphogenesis. Notably, maize has been a key system for delineating auxin biosynthetic pathways and offers many opportunities for future investigations on auxin metabolism. In addition, crosstalk between auxin and other phytohormones has been uncovered through gene expression studies and are important for leaf and root development in maize. Collectively these studies point to auxin as a cornerstone for maize biology that could be leveraged for improved crop resilience and yield."
Authors: Linfang Li and Xu Chen.
Research Square (2022)
Abstract: "Breeding crop varieties with high-yield and ideal plant architecture is a desirable goal of agricultural science. The success of ‘Green Revolution’ in cereal crops provides opportunities to incorporate phytohormones in crop breeding. Auxin is a critical phytohormone to determinate nearly all the aspects of plant development. Despite the current knowledge regarding auxin biosynthesis, auxin transport and auxin signaling has been well characterized in model Arabidopsis plants, how auxin regulates crop architecture is far from being understood and the introduction of auxin biology in crop breeding stays in the theoretical stage. Here, we give an overview on molecular mechanisms of auxin biology in Arabidopsis, and mainly summarize auxin contributions for crop plant development. Furthermore, we propose potential opportunities to integrate auxin biology in soybean breeding."
Authors: Shuifu Chen, Yuqun Huang, Jingluan Han, Shijuan Zhang, Qiaoyu Yang, Zhijie Li, Ya Zhang, Runyuan Mao, Ling Fan, Yaoguang Liu, Yuanling Chen and Xianrong Xie.
International Journal of Molecular Sciences (2022)
Abstract: "Tiller angle is an important trait that determines plant architecture and yield in cereal crops. Tiller angle is partially controlled during gravistimulation by the dynamic re-allocation of LAZY1 (LA1) protein between the nucleus and plasma membrane, but the underlying mechanism remains unclear. In this study, we identified and characterized a new allele of LA1 based on analysis of a rice (Oryza sativa L.) spreading-tiller mutant la1G74V, which harbors a non-synonymous mutation in the predicted transmembrane (TM) domain-encoding region of this gene. The mutation causes complete loss of shoot gravitropism, leading to prostrate growth of plants. Our results showed that LA1 localizes not only to the nucleus and plasma membrane but also to the endoplasmic reticulum. Removal of the TM domain in LA1 showed spreading-tiller phenotype of plants similar to la1G74V but did not affect the plasma membrane localization; thus, making it distinct from its ortholog ZmLA1 in Zea mays. Therefore, we propose that the TM domain is indispensable for the biological function of LA1, but this domain does not determine the localization of the protein to the plasma membrane. Our study provides new insights into the LA1-mediated regulation of shoot gravitropism."
Authors: Zhongqin Zhang, Le Gao, Meiyu Ke, Zhen Gao, Tianli Tu, Laimei Huang, Jiaomei Chen, Yuefeng Guan, Xi Huang and Xu Chen.
Journal of Integrative Plant Biology (2022)
Abstract: "Crop breeding during the Green Revolution resulted in high yields largely due to the creation of plants with semi-dwarf architectures that could tolerate high-density planting. Although semi-dwarf varieties have been developed in rice, wheat and maize, none was reported in soybean (Glycine max), and few genes controlling plant architecture have been characterized in soybean. Here, we demonstrate that the auxin efflux transporter PINFORMED1 (GmPIN1), which determines polar auxin transport, regulates the leaf petiole angle in soybean. CRISPR-Cas9-induced Gmpin1abc and Gmpin1bc multiple mutants displayed a compact architecture with a smaller petiole angle than wild-type plants. GmPIN1 transcripts and auxin were distributed asymmetrically in the petiole base, with high levels of GmPIN1a/c transcript and auxin in the lower cells, which resulted in asymmetric cell expansion. By contrast, the (iso)flavonoid content was greater in the upper petiole cells than in the lower cells. Our results suggest that (iso)flavonoids inhibit GmPIN1a/c expression to regulate the petiole angle. Overall, our study demonstrates that a signal cascade that integrates (iso)flavonoid biosynthesis, GmPIN1a/c expression, auxin accumulation, and cell expansion in an asymmetric manner creates a desirable petiole curvature in soybean. This study provides a genetic resource for improving soybean plant architecture."
Authors: Meiqing Xing, Wei Wang, Xing Fang and Hongwei Xue.
The Crop Journal (2022)
Abstract: "Leaf inclination, a component of crop architecture, influences photosynthetic efficiency and planting density. Various factors, particularly the phytohormones auxin and brassinosteroids (BRs), function in regulating lamina joint bending, and understanding of the genetic control of leaf inclination will help to elucidate the relevant regulatory network. Screening a rice T-DNA insertion population revealed a mutant that was insensitive to auxin and displayed an enlarged leaf angle due to increased cell length on the adaxial side of the lamina joint. Genetic analysis revealed that the increased leaf inclination was caused by T-DNA insertion in the promoter region of OsIAA6, resulting in elevated OsIAA6 expression. Further study showed that OsIAA6 interacts with OsARF1 to suppress auxin signaling and regulates leaf inclination. OsIAA6 mediates the BR effects on lamina joint development, and OsBZR1, the key transcription factor in BR signaling, binds directly to the promoter of OsIAA6 to stimulate its transcription. These results indicate the roles of the OsIAA6–OsARF1 module in regulating rice leaf inclination and suggest the synergistic effects of the phytohormones auxin and BR."
Authors: Hyodong Lee, Anindya Ganguly, Song Baik and Hyung-Taeg Cho.
The Plant Cell (2021)
Abstract: "PIN-FORMED (PIN)-mediated polar auxin transport is involved in key developmental processes in plants. Various internal and external cues influence plant development via the modulation of intracellular PIN polarity and, thus, the direction of polar auxin transport, but the mechanisms underlying these processes remain largely unknown. PIN proteins harbor a hydrophilic loop (HL) that has important regulatory functions; here, we used the HL as bait in protein pulldown screening for modulators of intracellular PIN trafficking in Arabidopsis thaliana. CPK29, a Ca2+-dependent protein kinase, was identified and shown to phosphorylate specific target residues on the PIN-HL that were not phosphorylated by other kinases. Furthermore, loss of CPK29 or mutations of the phospho-target residues in PIN-HLs significantly compromised intracellular PIN trafficking and polarity, causing defects in PIN-mediated auxin redistribution and biological processes such as lateral root formation, root twisting, hypocotyl gravitropism, phyllotaxis, and reproductive development. These findings indicate that CPK29 directly interprets Ca2+ signals from internal and external triggers, resulting in the modulation of PIN trafficking and auxin responses."
Authors: Shiqing Dong, Xianxin Dong, Xiaokang Han, Fan Zhang, Yu Zhu, Xiaoyun Xin, Ying Wang, Yuanyi Hu, Dingyang Yuan, Jianping Wang, Zhou Huang, Fuan Niu, Zejun Hu, Peiwen Yan, Liming Cao, Haohua He, Junru Fu, Yeyun Xin, Yanning Tan, Bigang Mao, Bingran Zhao, Jinshui Yang, Longping Yuan and Xiaojin Luo.
PNAS (2021)
Significance: Rice breeding programs aim to develop cultivars with improved traits, including high grain yield and superior quality. In rice, OsPDCD5 encodes a programmed cell death 5 protein. Targeted mutagenesis of OsPDCD5 enhanced grain yield and plant architecture. Statistical analysis indicated that plot grain yield of OsPDCD5 knockout lines was enhanced by 6.25 to 20.13% in 11 popular or newly bred rice cultivars compared with the corresponding wild types. The OsPDCD5 knockout lines showed increases in milled rice percentage and gel consistency, and a decrease in amylose content. Our results provide insight into the molecular mechanism by which OsPDCD5 influences grain yield and plant architecture, and highlight a promising candidate gene for use in breeding programs designed to develop super rice cultivars.
Abstract: "Plant architecture is an important agronomic trait that affects crop yield. Here, we report that a gene involved in programmed cell death, OsPDCD5, negatively regulates plant architecture and grain yield in rice. We used the CRISPR/Cas9 system to introduce loss-of-function mutations into OsPDCD5 in 11 rice cultivars. Targeted mutagenesis of OsPDCD5 enhanced grain yield and improved plant architecture by increasing plant height and optimizing panicle type and grain shape. Transcriptome analysis showed that OsPDCD5 knockout affected auxin biosynthesis, as well as the gibberellin and cytokinin biosynthesis and signaling pathways. OsPDCD5 interacted directly with OsAGAP, and OsAGAP positively regulated plant architecture and grain yield in rice. Collectively, these findings demonstrate that OsPDCD5 is a promising candidate gene for breeding super rice cultivars with increased yield potential and superior quality."
|
Authors: Yuqi Liu, Shangyu Chen, Sikander Pal, Jingquan Yu, Yanhong Zhou, Lam-Son Phan Tran and Xiaojian Xia.
New Crops (2024)
Abstract: "Plants have evolved varied structures for environmental adaptation. Shoot branching, as a part of plant architecture, influences the allocation of sugars produced by photosynthesis and thus greatly impacts crop yields. The activity of axillary meristem, and apical dominance governs the shoot branching patterns. In this review, we summarize the key factors involved in the formation of lateral branches, and the mechanisms of how these factors are interconnected. In particular, we focus on recent advances in understanding how sugar and environmental signals affect the hormonal signaling network to regulate apical dominance. Ultimately, we propose that epigenetic modifications are critical mechanisms underlying the plasticity of shoot branching, and that precise targeted gene editing is promising for shaping the ideal plant architecture."
Authors: Connor Tansley, Nicola J. Patron, and Sarah Guiziou
ACS Synthetic Biology (2024)
Abstract: "Many plant species are grown to enable access to specific organs or tissues, such as seeds, fruits, or stems. In some cases, a value is associated with a molecule that accumulates in a single type of cell. Domestication and subsequent breeding have often increased the yields of these target products by increasing the size, number, and quality of harvested organs and tissues but also via changes to overall plant growth architecture to suit large-scale cultivation. Many of the mutations that underlie these changes have been identified in key regulators of cellular identity and function. As key determinants of yield, these regulators are key targets for synthetic biology approaches to engineer new forms and functions. However, our understanding of many plant developmental programs and cell-type specific functions is still incomplete. In this Perspective, we discuss how advances in cellular genomics together with synthetic biology tools such as biosensors and DNA-recording devices are advancing our understanding of cell-specific programs and cell fates. We then discuss advances and emerging opportunities for cell-type-specific engineering to optimize plant morphology, responses to the environment, and the production of valuable compounds."
Authors: Guan-Ting Erica Chen, Jian You Wang, Cristina Votta, Justine Braguy, Muhammad Jamil, Gwendolyn K. Kirschner, Valentina Fiorilli, Lamis Berqdar, Aparna Balakrishna, Ikram Blilou, Luisa Lanfranco and Salim Al-Babili.
PNAS (2023)
Significance: Strigolactones (SLs) are multifunctional, structurally diverse secondary metabolites fulfilling the function of a hormone. Whether a particular SL exerts specific functions is one of the most important questions in SL biology. Here, we generated and characterized rice mutants lacking the common SLs 4-deoxyorobanchol and/or its derivative orobanchol, which represent one of the two SL subfamilies, i.e., canonical SLs. We show that 4-deoxyorobanchol is not a determinant of shoot branching, but has a specific function as a regulator of shoot, root, and panicle growth. Accumulation of 4-deoxyorobanchol affects auxin homeostasis and negatively impacts the symbiosis with mycorrhizal fungi. Our data reveal specific hormonal functions of canonical SLs and pave the way for targeted modulation of rice architecture and rhizospheric interactions.
Abstract: "Strigolactones (SLs) regulate many developmental processes, including shoot-branching/tillering, and mediate rhizospheric interactions. SLs originate from carlactone (CL) and are structurally diverse, divided into a canonical and a noncanonical subfamily. Rice contains two canonical SLs, 4-deoxyorobanchol (4DO) and orobanchol (Oro), which are common in different plant species. The cytochrome P450 OsMAX1-900 forms 4DO from CL through repeated oxygenation and ring closure, while the homologous enzyme OsMAX1-1400 hydroxylates 4DO into Oro. To better understand the biological function of 4DO and Oro, we generated CRISPR/Cas9 mutants disrupted in OsMAX1-1400 or in both OsMAX1-900 and OsMAX1-1400. The loss of OsMAX1-1400 activity led to a complete lack of Oro and an accumulation of its precursor 4DO. Moreover, Os1400 mutants showed shorter plant height, panicle and panicle base length, but no tillering phenotype. Hormone quantification and transcriptome analysis of Os1400 mutants revealed elevated auxin levels and changes in the expression of auxin-related, as well as of SL biosynthetic genes. Interestingly, the Os900/1400 double mutant lacking both Oro and 4DO did not show the observed Os1400 architectural phenotypes, indicating their being a result of 4DO accumulation. Treatment of wild-type plants with 4DO confirmed this assumption. A comparison of the Striga seed germinating activity and the mycorrhization of Os900, Os900/1400, and Os1400 loss-of-function mutants demonstrated that the germination activity positively correlates with 4DO content while disrupting OsMAX1-1400 has a negative impact on mycorrhizal symbiosis. Taken together, our paper deciphers the biological function of canonical SLs in rice and reveals their particular contributions to establishing architecture and rhizospheric communications."
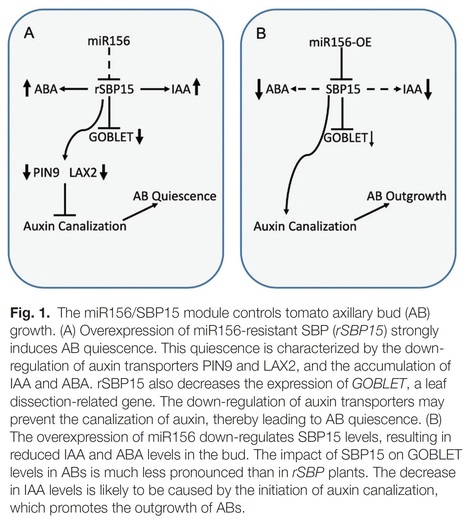
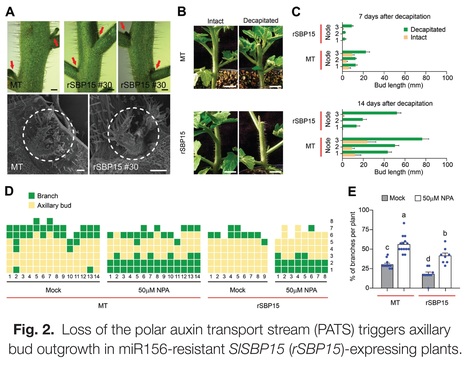
Authors: Carlos Hernán Barrera-Rojas, Mateus Henrique Vicente, Diego Armando Pinheiro Brito, Eder M. Silva, Aitor Muñoz Lopez, Leticia F. Ferigolo, Rafael Monteiro do Carmo, Carolina M. S. Silva, Geraldo F. F. Silva, Joao P. O. Correa, Marcela M. Notini, Luciano Freschi, Pilar Cubas and Fabio T. S. Nogueira.
Journal of Experimental Botany (2023)
Abstract: "The miRNA156 (miR156)/SQUAMOSA PROMOTER-BINDING PROTEIN-LIKE (SPL/SBP) regulatory hub is highly conserved among phylogenetically distinct species, but how it interconnects multiple pathways to converge to common integrators controlling shoot architecture is still unclear. Here, we demonstrated that the miR156/SlSBP15 node modulates tomato shoot branching by connecting multiple phytohormones with classical genetic pathways regulating both axillary bud development and outgrowth. miR156-overexpressing plants (156-OE) displayed high shoot branching, whereas plants overexpressing a miR156-resistant SlSBP15 allele (rSBP15) showed arrested shoot branching. Importantly, the rSBP15 allele was able to partially restore the wild-type shoot branching phenotype in the 156-OE background. rSBP15 plants have tiny axillary buds, and their activation is dependent on shoot apex-derived auxin transport inhibition. Hormonal measurements revealed that indole-3-acetic acid (IAA) and abscisic acid (ABA) concentrations were lower in 156-OE and higher in rSBP15 axillary buds, respectively. Genetic and molecular data indicated that SlSBP15 regulates axillary bud development and outgrowth by inhibiting auxin transport and GOBLET (GOB) activity, and by interacting with tomato BRANCHED1b (SlBRC1b) to control ABA levels within axillary buds. Collectively, our data provide a new mechanism by which the miR156/SPL/SBP hub regulates shoot branching, and suggest that modulating SlSBP15 activity might have potential applications in shaping tomato shoot architecture."
Authors: Hongxiang Lou, Bowen Zhao, Yan Peng, Ali Mahmoud El-Badri, Maria Batool, Chunyun Wang, Zongkai Wang, Wei Huang, Tianyao Wang, Zhen Li, Zhenghua Xu, Jing Wang, Bo Wang, Jie Kuai and Guangsheng Zhou.
Field Crops Research (2023)
Highlights • Various plant architecture rapeseeds responded differently to nitrogen and density. • HS5+/sca was suitable for reducing nitrogen rate and increasing plant density. • BnaA3. IAA7 mutation caused more lateral roots and root biomass in HS5+/sca. • Auxin level and its signaling genes expression affected root-crown growth.
Abstract: "Context - With the increasing global food and oil security problems, improvement in rapeseed (Brassica napus L.) yield become urgent. Moreover, previous studies rarely used the semi-dwarf and compact plant type of rapeseed to improve yield potential. Objective - Herein, we aimed to investigate the effects of different nitrogen application rates and plant density on root growth and yield-related traits using three genotypes with different plant types to clarify various regulatory mechanisms using semi-dwarf and compact plant type mutants to achieve high yield. Methods - Split-split-plot experiments with three nitrogen application rates (N1: N2: N3, 120: 240: 360 kg ha−1) and three plant densities (D1: D2: D3, 15 × 104: 45 × 104: 75 × 104 plants·ha−1) were conducted during the three growing seasons (2019–2022) using semi-dwarf mutant HS5sca, and HS5 (wild type) as well as their F1 hybrid HS5+/sca. Results - With decreasing nitrogen application rate (N3 to N2), the expression level of IAA signal response genes BnaA07. GH3 and BnaA03. IAA13 was decreased in root, while increased in shoot at the early-flowering stage. Moreover, IAA content was decreased in roots and aboveground parts, whereas soluble sugar content of root bleeding sap was increased and volume of root bleeding sap was decreased in three genotypes at early-flowering stage. Additionally, nitrogen accumulation per plant and yield per plant was decreased, while population nitrogen accumulation and population yield were increased with decreasing nitrogen application rate in three studied genotypes. On the other side, with increasing plant density, the expression of BnaA07. GH3 and BnaA03. IAA13 genes were first increased and then decreased in root tissues. In addition, root surface area, the total volume of root bleeding sap, and nitrogen accumulation were decreased, which decreased dry matter and yield per plant. However, total root area, population nitrogen accumulation, and population yield in the three genotypes were increased, and yield was the highest at D2. The population yield of HS5sca, HS5+/sca, and HS5 achieved the largest increase of 26.0%, 10.3%, and 9.5% under N2D2 in 2019, 6.9%, 8.4%, and 4.7% under N3D2 in 2020, respectively, as compared to N3D1. Moreover, the morphological indices of HS5+/sca were greater than other genotypes, with the highest population yield. Conclusions - Appropriate nitrogen application rate with plant density optimized the root architecture and aboveground type of HS5+/sca that improved auxin level through affecting the downstream auxin-related genes, as well as improved dry matter accumulation and distribution, which increased yield potential. Compared with the two parents, F1 genotype HS5+/sca was suitable for nitrogen fertilizer (240 kg ha−1) and plant density (45–75 × 104 plants ha−1). Implications - + /sca heterozygous genotypes might promote root growth under low nitrogen application rate and high plant density. A strong root system is an important guarantee of high yield; thus, this heterozygous site could use to introduce excellent hybrids in rapeseed production to further improve root traits and optimize cultivation factors to increase seed yield.
Authors: Tengfei Liu, Liepeng Dong, Enshuang Wang, Shengxuan Liu, Yunxia Cheng, Ji Zhao, Shijing Xu, Zhen Liang, Hui Ma, Bihua Nie and Botao Song.
Journal of Experimental Botany (2023)
Abstract: "Abscisic acid (ABA) is critical in drought tolerance and plant growth. Group A protein type 2C phosphatases (PP2Cs) are negative regulators of ABA signaling and plant adaptation to stress. However, our knowledge about the functions of potato group A PP2Cs is limited. Here, we report that potato group A PP2C StHAB1 is broadly expressed in potato plants and strongly induced by ABA and drought. Suppression of StHAB1 enhanced potato ABA sensitivity and drought tolerance, whereas overexpression of the dominant mutant StHAB1G276D compromised ABA sensitivity and drought tolerance. StHAB1 interacts with almost all ABA receptors and the Snf1-Related Kinase OST1. Suppressing StHAB1 and overexpressing StHAB1G276D alter potato growth morphology; notably, overexpression of StHAB1G276D causes excessive shoot branching in potato. RNA-seq analyses identified that auxin efflux carrier genes, StPIN3, StPIN5, and StPIN8, were upregulated in StHAB1G276D-overexpressed axillary buds. Correspondingly, auxin concentration was reduced in StHAB1G276D-overexpressed axillary buds, consistent with the repressing role of auxin in lateral branch outgrowth. The expression of BRANCHED1s (StBRC1a and StBRC1b) does not change in StHAB1G276D-overexpressed axillary buds, suggesting that overexpression of StHAB1G276D caused axillary bud outgrowth is not due to regulating BRC1 expression. Our findings demonstrate that StHAB1 is vital in potato drought tolerance and shoot branching."
Authors: Lei Zhao, Yueting Zheng, Ying Wang, Shasha Wang, Tongzhu Wang, Canguan Wang, Yue Chen, Kunpu Zhang, Ning Zhang, Zhongdong Dong and Feng Chen.
Plant Biotechnology Journal (2022)
Abstract: "Tiller angle is one of the most important agronomic traits and one key factor for wheat ideal plant architecture, which can both increase photosynthetic efficiency and greatly enhance grain yield. Here, a deacetylase HST1-like (TaHST1L) gene controlling wheat tiller angle was identified by the combination of a genome-wide association study (GWAS) and bulked segregant analysis (BSA). Ethyl methane sulfonate (EMS)-mutagenized tetraploid wheat lines with the premature stop codon of TaHST1L exhibited significantly smaller tiller angles than the wild type. TaHST1L-overexpressing (OE) plants exhibited significantly larger tiller angles and increased tiller numbers in both winter and spring wheat, while TaHST1L-silenced RNAi plants displayed significantly smaller tiller angles and decreased tiller numbers. Moreover, TaHST1L strongly interacted with TaIAA17 and inhibited its expression at the protein level, and thus possibly improved the content of endogenous auxin in the basal tissue of tillers. The transcriptomics and metabolomics results indicated that TaHST1L might change plant architecture by mediating auxin signal transduction and regulating endogenous auxin levels. In addition, a 242-bp insertion/deletion (InDel) in the TaHST1L-A1 promoter altered transcriptional activity and TaHST1L-A1b allele with the 242-bp insertion widened the tiller angle of TaHST1L-OE transgenic rice plants. Wheat varieties with TaHST1L-A1b allele possessed the increased tiller angle and grain yield. Further analysis in wheat and its progenitors indicated that the 242-bp InDel possibly originated from wild emmer and was strongly domesticated in the current varieties. Therefore, TaHST1L involved in the auxin signaling pathway showed the big potential to improve wheat yield by controlling plant architecture."
Authors: Yan Li, Yizhou He, Zhixin Liu, Tian Qin, Lei Wang, Zhihui Chen, Biaoming Zhang, Haitao Zhang, Haitao Li, Li Liu, Jian Zhang and Wenya Yuan.
The Plant Journal (2022)
Abstract: "As a multigenic trait, rice tillering can optimize plant architecture for the maximum agronomic yield. SQUAMOSA PROMOTER BINDING PROTEIN-LIKE14 (OsSPL14) has been demonstrated to be necessary and sufficient to inhibit rice branching, but the underlying mechanism remains largely unclear. Here, we demonstrated that OsSPL14, which is cleaved by miR529 and miR156, inhibits tillering by fine-tuning auxin transport in rice. RNA interference of OsSPL14 (OsSPL14 RNAi) or miR529 and miR156 overexpression significantly increased the tiller number, whereas OsSPL14 overexpression obviously decreased the tiller number. Histological analysis revealed that the OsSPL14-overexpressing line had normal initiation of axillary buds but inhibited outgrowth of tillers. Moreover, OsSPL14 was found to be responsive to indole-acetic acid (IAA) and 1-naphthylphthalamic acid (NPA), and OsSPL14 RNAi reduced polar auxin transport (PAT) and increased NPA sensitivity of rice plants. Further analysis revealed that OsSPL14 directly binds to the promoter of PIN-FORMED 1b (OsPIN1b) and PIN-LIKE6b (PILS6b) to positively regulate their expression. OsPIN1b and PILS6b were highly expressed in axillary buds and proved to be involved in bud outgrowth. Loss of function of OsPIN1b or PILS6b increased the tiller number of rice. Taken together, our findings suggested that OsSPL14 can control axillary bud outgrowth and tiller number by activating the expression of OsPIN1b and PILS6b to fine-tune auxin transport in rice."
Authors: Fu LI, Dong YAN, Li-feng GAO, Pan LIU, Guang-yao ZHAO, Ji-zeng JIA and Zheng-long REN.
Journal of Integrative Agriculture (2022)
Abstract: "Bread wheat (Triticum aestivum L.) is one of the most important staple crops worldwide. The phytohormone auxin plays critical roles in the regulation of plant growth and development. However, only a few auxin-related genes have been genetically demonstrated to be involved in the control of plant architecture in wheat thus far. In this study, we characterized an auxin-related gene in wheat, TaIAA15, and found that its ectopic expression in rice decreased the plant height and increased the leaf angle. Correlation analysis indicated that TaIAA15-3B was associated with plant height (Ph), spike length (SL) and 1 000-grain weight (TGW) in wheat, and Hap-II of TaIAA15-3B was the most favored allele and selected by modern breeding in China. This study sheds light on the role of auxin signaling on wheat plant architecture as well as yield related traits."
Authors: Chaohui Ding, Xianhui Lin, Ying Zuo, Zhilin Yu, Scott R. Baerson, Zhiqiang Pan, Rensen Zeng and Yuanyuan Song.
The Plant Journal (2021)
Abstract: "Tiller angle is an important determinant of plant architecture in rice (Oryza sativa L.). Auxins play a critical role in determining plant architecture; however, the underlying metabolic and signaling mechanisms are still largely unknown. In this study, we have identified a member of the bZIP family of TGA class transcription factors, OsbZIP49, that participates in the regulation of plant architecture and is specifically expressed in gravity-sensing tissues, including the shoot base, nodes and lamina joints. Transgenic rice plants overexpressing OsbZIP49 displayed a tiller-spreading phenotype with reduced plant height and internode lengths. In contrast, CRISPR/Cas9-mediated knockout of OsbZIP49 resulted in a compact architecture. Follow-up studies indicated that the effects of OsbZIP49 on tiller angles are mediated through changes in shoot gravitropic responses. Additionally, we provide evidence that OsbZIP49 activates the expression of indole-3-acetic acid-amido synthetases OsGH3-2 and OsGH3-13 by directly binding to TGACG motifs located within the promoters of both genes. Increased GH3-catalyzed conjugation of indole-3-acetic acid (IAA) in rice transformants overexpressing OsbZIP49 resulted in the increased accumulation of IAA-Asp and IAA-Glu, and a reduction in local free auxin, tryptamine and IAA-Glc levels. Exogenous IAA or naphthylacetic acid (NAA) partially restored shoot gravitropic responses in OsbZIP49-overexpressing plants. Knockout of OsbZIP49 led to reduced expression of both OsGH3-2 and OsGH3-13 within the shoot base, and increased accumulation of IAA and increased OsIAA20 expression levels were observed in transformants following gravistimulation. Taken together, the present results reveal the role transcription factor OsbZIP49 plays in determining plant architecture, primarily due to its influence on local auxin homeostasis."
Authors: Udita Basu and Swarup K. Parida.
Plant Biotechnology Journal (2021)
Abstract: "Plants have adapted to different environmental niches by fine-tuning the developmental factors working together to regulate traits. Variations in the developmental factors result in a wide range of quantitative variations in these traits that helped plants survive better. The major developmental pathways affecting plant architecture are also under the control of such pathways. Most notable are the CLAVATA-WUSCHEL pathway regulating shoot apical meristem fate, GID1-DELLA module influencing plant height and tillering, LAZY1-TAC1 module controlling branch/tiller angle, and the TFL1-FT determining the floral fate in plants. Allelic variants of these key regulators selected during domestication shaped the crops the way we know them today. There is immense yield potential in the “ideal plant architecture” of a crop. With the available genome editing techniques, possibilities are not restricted to naturally occurring variations. Using a transient reprogramming system, one can screen the effect of several developmental gene expressions in novel ecosystems to identify the best targets. We can use the plant's fine-tuning mechanism for customizing crops to specific environments. The process of crop domestication can be accelerated with a proper understanding of these developmental pathways. It is time to step forward towards the next-generation molecular breeding for restructuring plant types in crops ensuring yield stability."
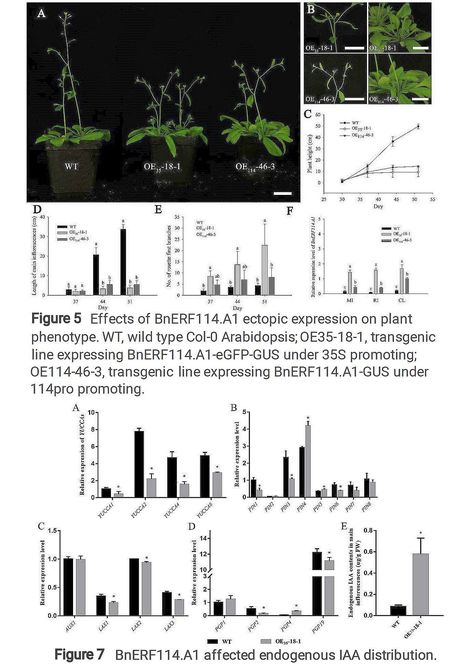
Authors: Jinyang Lyu, Yuan Guo, Chunlei Du, Haibo Yu, Lijian Guo, Li Liu, Xinfa Wang, Huixian Zhao and Shengwu Hu.
Research Square (2021)
Abstract: "Plant architecture is very important for rapeseed breeding. Here, we reported an ETHYLENE RESPONSE FACTOR (ERF) transcription factor BnERF114.A1 of Brassica napus participating in plant architecture regulation. BnERF114.A1 is a member of ERF family group x-a, encoding a putative protein of 252 aa which consisting of an AP2/ERF domain and a conserved CMX-1 motif. BnERF114.A1 located in nucleus and had transcriptional activity with its functional region located in 142 aa ~ 252 aa of its C-terminus. The GUS staining analysis revealed that BnERF114.A1 highly expressed in leaf primordia, shoot apical meristem, leaf marginal meristem, and reproductive organs. Ectopic expression of BnERF114.A1 in Arabidopsis reduced plant height, increased branch numbers and silique numbers per plants, and finally increased seed yield per plant. Further investigation demonstrated that overexpression of BnERF114.A1 can inhibit IAA efflux and cause accumulation of auxin in apex, and arrest apical dominance in Arabidopsis. The findings suggested BnERF114.A1 could provide a candidate gene for rapeseed plant architecture molecular breeding."
|


 Your new post is loading...
Your new post is loading...