Research and publish the best content.
Get Started for FREE
Sign up with Facebook Sign up with X
I don't have a Facebook or a X account
Already have an account: Login
 Your new post is loading... Your new post is loading...
 Your new post is loading... Your new post is loading...
|
|




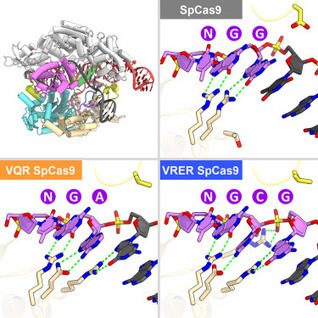

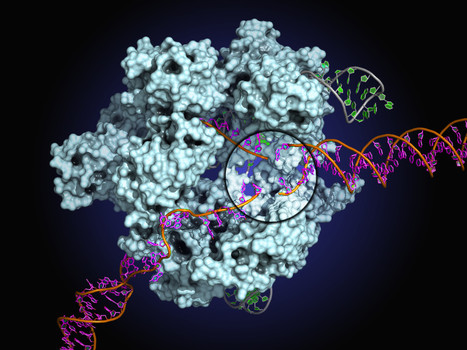





In this report, the scientists present the high-resolution crystal structures of the three SpCas9 variants in complexes with a single-guide RNA and its altered PAM-containing, partially double-stranded DNA targets. A structural comparison of the three SpCas9 variants with wild-type SpCas9 revealed that the multiple mutations synergistically induce an unexpected displacement in the phosphodiester backbone of the PAM duplex, thereby allowing the SpCas9 variants to directly recognize the altered PAM nucleotides. These findings explain the altered PAM specificities of the SpCas9 variants and establish a framework for further rational engineering of CRISPR-Cas9.