Research and publish the best content.
Get Started for FREE
Sign up with Facebook Sign up with X
I don't have a Facebook or a X account
Already have an account: Login
 Your new post is loading... Your new post is loading...
 Your new post is loading... Your new post is loading...
|
|







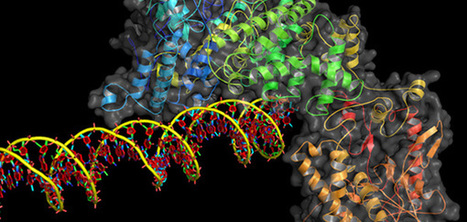
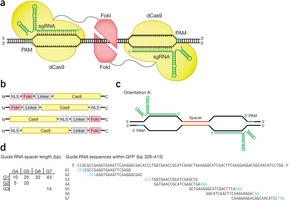





The scientists recently reported Digenome-seq (digested genome sequencing), a method for in vitro identification of potential off-target sites, and They evaluated the specificity of CRISPR–Cas9 and CRISPR–Cpf1 endonuclease by whole-genome sequencing. Digenome-seq pinpoints the exact location of double-strand break (DSB) sites by recognizing specific patterns of aligned reads. However, the analysis pipeline presented in their previous report required extensive manual interaction and produced several large intermediate files, resulting in a long running time. Here, the scientists present a redesigned analysis tool for Digenome-seq data that runs on web browsers. The core algorithm of the tool is written in C++ and compiled to asm.js (http://asmjs.org/), a preoptimized subset of JavaScript. Users can instantly perform the complete analysis in an ordinary web browser (Supplementary Note 1) with fast execution speed without uploading any data to a server and without local tool installation. In their benchmark, the full analysis for 100 GB of BAM file took 3 h for whole analysis on Intel i5 3570k central processing unit in a single thread.