Research and publish the best content.
Get Started for FREE
Sign up with Facebook Sign up with X
I don't have a Facebook or a X account
Already have an account: Login
 Your new post is loading... Your new post is loading...
 Your new post is loading... Your new post is loading...
|
|









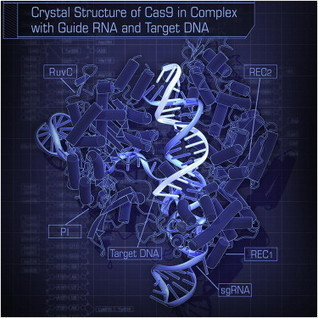






Here the authors demonstrate that the class 2 type VI RNA-guided RNA-targeting CRISPR–Cas effector Cas13a (previously known as C2c2) can be engineered for mammalian cell RNA knockdown and binding. Their results establish CRISPR–Cas13a as a flexible platform for studying RNA in mammalian cells and therapeutic development.