Ni OGM, ni processus classique de sélection par croisements successifs: des chercheurs ont dévoilé dimanche une technique révolutionnaire pour produire en un temps record des semences à fort rendement ou résistantes au changement climatique.
Research and publish the best content.
Get Started for FREE
Sign up with Facebook Sign up with X
I don't have a Facebook or a X account
Already have an account: Login

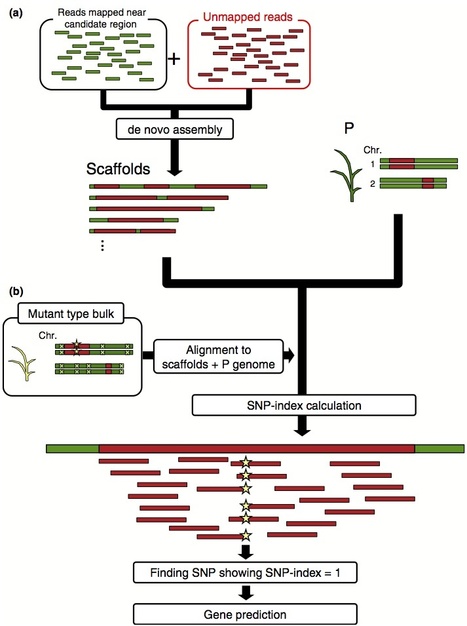
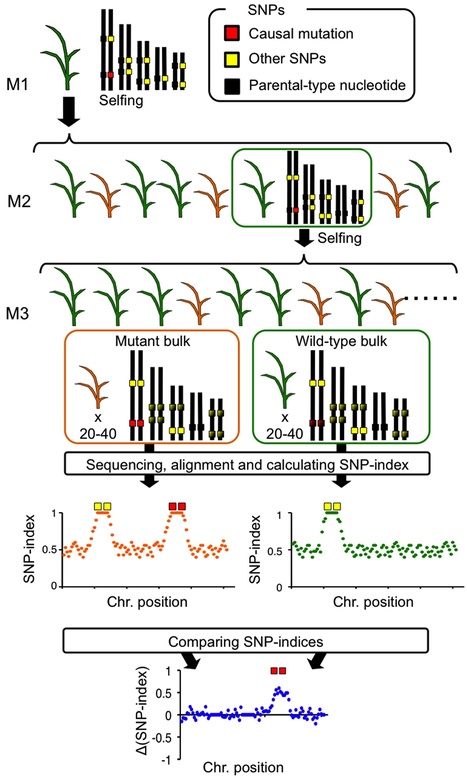
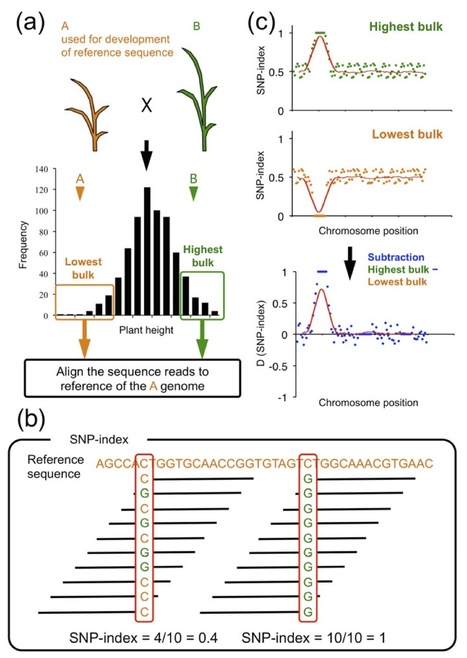
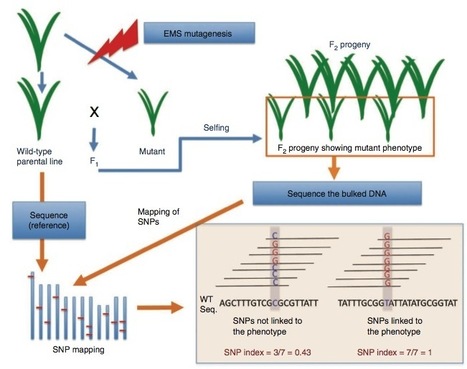
Everything about MutMap, a method based on whole-genome sequencing of pooled DNA from a segregating population http://tinyurl.com/mutmap-nbt
Curated by
Kamoun Lab @ TSL
 Your new post is loading... Your new post is loading...
 Your new post is loading... Your new post is loading...
|
|