 Your new post is loading...
 Your new post is loading...

|
Scooped by
dromius
|
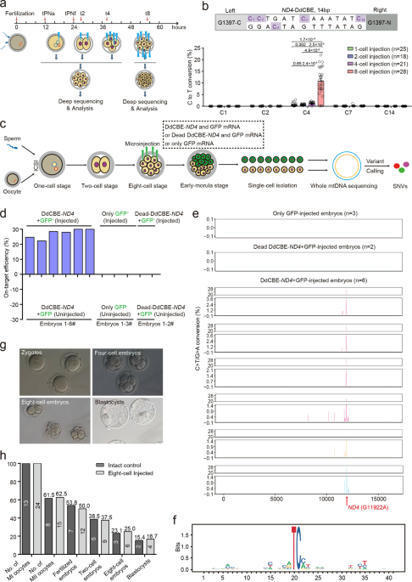
(via T. Schreiber, thx) Wei et al, 2022 Although ZFN and TALEN have been previously engineered to successfully cut and eliminate mtDNA in a programmable way5,6,7,8,9, correction of disease-causing point mutations in mtDNA remained challenging. Recently, DddA-derived cytosine base editors (DdCBEs) have been developed to specifically induce C-to-T conversion in mtDNA by fusion of sequence-programmable TALE and split deaminase derived from interbacterial toxins10.
In this study, we sought to test DdCBE’s ability for base editing of mtDNA in human embryos.
Taken together, our results demonstrate that DdCBE is an effective base editor for inducing point mutations in mtDNA of human embryos, and the efficiency is much higher in 8-cell embryos. Our 8-cell injection method could help generate mitochondrial disease models as well as derived embryonic stem cells for functional investigation of disease-associated mutations in mtDNA.

|
Scooped by
dromius
|
Liu, et al 2021
Engineered transcriptional activator-like effectors (TALEs) are versatile tools for genome manipulation with applications in research and clinical contexts. One current drawback of TALEs is that the 5′ nucleotide of the target is specific for thymine (T). TALE domains with alternative 5′ nucleotide specificities could expand the scope of DNA target sequences that can be bound by TALEs. Another drawback of TALEs is their tendency to bind and cleave off-target sequence, which hampers their clinical application and renders applications requiring high-fidelity binding unfeasible. This disclosure provides methods and strategies for the continuous evolution of proteins comprising DNA-binding domains, e.g., TALE domains. In some aspects, this disclosure provides methods and strategies for evolving such proteins under positive selection for a desired DNA-binding activity and/or under negative selection against one or more undesired (e.g., off-target) DNA-binding activities. Some aspects of this disclosure provide engineered TALE domains and TALEs comprising such engineered domains, e.g., TALE nucleases (TALENs), TALE transcriptional activators, TALE transcriptional repressors, and TALE epigenetic modification enzymes, with altered 5′ nucleotide specificities of target sequences. Engineered TALEs that target ATM with greater specificity are also provided.

|
Scooped by
dromius
|
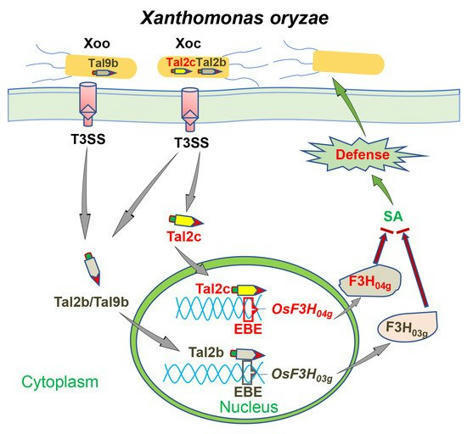
(via T. Schreiber, thx) Wu et al, 2021 Xanthomonas oryzae delivers transcription activator-like effectors (TALEs) into plant cells to facilitate infection. Following economic principles, the redundant TALEs are rarely identified in Xanthomonas. Previously, we identified the Tal2b, which activates the expression of the rice 2-oxoglutarate-dependent dioxygenase gene OsF3H03g to promote infection in the highly virulent strain of X. oryzae pv. oryzicola HGA4. Here, we reveal that another clustered TALE, Tal2c, also functioned as a virulence factor to target rice OsF3H04g, a homologue of OsF3H03g. Transferring Tal2c into RS105 induced expression of OsF3H04g to coincide with increased susceptibility in rice. Overexpressing OsF3H04g caused higher susceptibility and less salicylic acid (SA) production compared to wild-type plants. Moreover, CRISPR–Cas9 system-mediated editing of the effector-binding element in the promoters of OsF3H03g or OsF3H04g was found to specifically enhance resistance to Tal2b- or Tal2c-transferring strains, but had no effect on resistance to either RS105 or HGA4. Furthermore, transcriptome analysis revealed that several reported SA-related and defense-related genes commonly altered expression in OsF3H04g overexpression line compared with those identified in OsF3H03g overexpression line. Overall, our results reveal a functional redundancy mechanism of pathogenic virulence in Xoc in which tandem Tal2b and Tal2c specifically target homologues of host genes to interfere with rice immunity by reducing SA.

|
Scooped by
dromius
|
(via T. Schreiber, thx) Wu et al, 2021 Transcription activator-like (TAL) effectors are major virulence factors secreted by the type III secretion systems of Xanthomonas oryzae pv. oryzicola (Xoc) and X. oryzae pv. oryzae (Xoo), causing bacterial leaf streak and bacterial blight, respectively, in rice. However, the knowledge of Xoc TAL effector function in promoting bacterial virulence remains limited.
Here, we isolated the highly virulent Xoc strain HGA4 from the outbreak region of Huanggang (Hubei, China), which contains four TAL effectors not found in the Chinese model strain RS105. Among these, Tal2b was selected for introduction into RS105, which resulted in a longer lesion length than that in the control.
Tal2b directly binds to the promoter region of the gene and activates the expression of OsF3H03g, which encodes 2-oxoglutarate-dependent dioxygenase in rice. OsF3H03g negatively regulates salicylic acid (SA)-related defense by directly reducing SA, and it plays a positive role in susceptibility to both Xoc and Xoo in rice.
OsF3H03g interacts with a uridine diphosphate-glycosyltransferase protein (OsUGT74H4), which positively regulates bacterial leaf streak susceptibility and may inactivate SA via glycosylation modification.

|
Scooped by
dromius
|
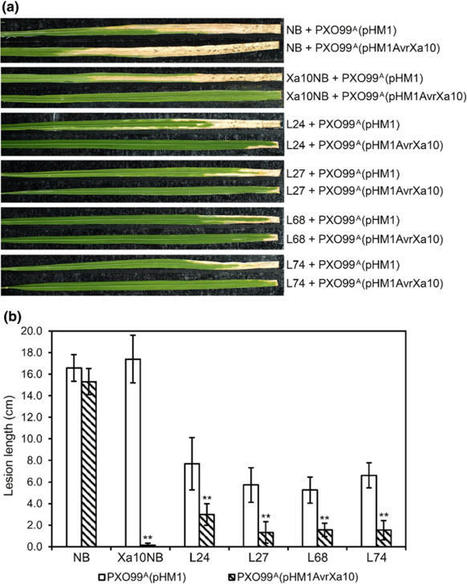
(via T. Schreiber, thx!) Gui et al, 2021 The hypersensitive response (HR) is a form of programmed cell death of plant cells occurring in the local region surrounding pathogen infection site to prevent the spread of infection by pathogens. Bax, a mammalian pro-apoptotic member of Bcl-2 family, triggers HR-like cell death when expressed in plants. However, constitutive expression of the Bax gene negatively affects plant growth and development. The Xa10 gene in rice (Oryza sativa) is an executor resistance (R) gene that confers race-specific disease resistance to Xanthomonas oryzae pv. oryzae strains harboring TAL effector gene AvrXa10. In this study, the Xa10 promoter was used to regulate heterologous expression of the Bax gene from mouse (Mus musculus) in Nicotiana benthamiana and rice. Cell death was induced in N. benthamiana after co-infiltration with the PXa10:Bax:TXa10 gene and the PPR1:AvrXa10:TNos gene. Transgenic rice plants carrying the PXa10:Bax:TXa10 gene conferred specific disease resistance to Xa10-incompatible X. oryzae pv. oryzae strain PXO99A(pHM1AvrXa10), but not to the Xa10-compatible strain PXO99A(pHM1). The resistance specificity was confirmed by the AvrXa10-dependent induction of the PXa10:Bax:TXa10 gene in transgenic rice. Our results demonstrated that the inducible expression of the Bax gene in transgenic rice was achieved through the control of the executor R gene promoter and the heterologous expression of the pro-apoptosis regulator gene in rice conferred disease resistance to X. oryzae pv. oryzae.

|
Scooped by
dromius
|
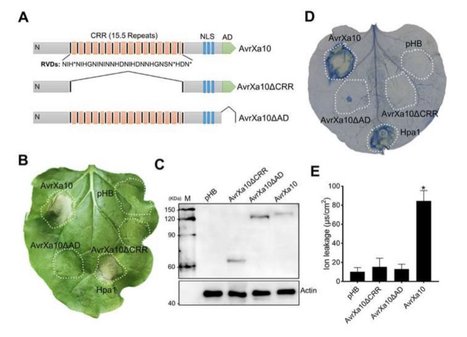
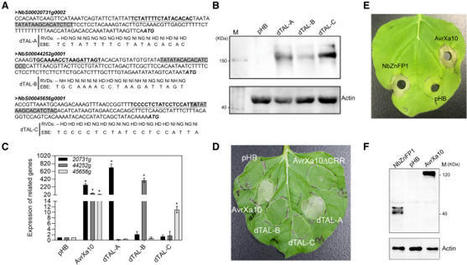
Haq et al, 2021 Xanthomonas oryzae pv. oryzae (Xoo), the causal agent of bacterial leaf blight (BLB) in rice, delivers transcription activator-like effector (TALE) proteins into host cells to activate susceptibility (S) or resistance (R) genes for disease or immunity, respectively. Nonhost plants serve as potential reservoirs of R genes; consequently, nonhost R genes may trap TALEs to trigger an immune response. In this study, we screened 17 Xoo TALEs for their ability to induce a hypersensitive response (HR) in the nonhost Nicotiana benthamiana (Nb); only AvrXa10 elicited a HR when transiently expressed in Nb. The HR generated by AvrXa10 required both the central repeat region and activation domain, suggesting specific interaction of AvrXa10 with a potential R-like gene in nonhost plant. Evans blue staining and ion leakage measurements confirmed that the AvrXa10-triggered HR was a form of cell death, and transient expression of AvrXa10 in Nb induced immune responses. Based on transcriptome profiling and prediction of effector binding sites, genes targeted by AvrXa10 in the Nb genome were identified. Using several approaches (in vivo reporter assays, electrophoretic mobility shift assays, targeted designer TALEs and on-spot gene silencing), we confirmed that AvrXa10 targets NbZnFP1, a C2H2-type zinc finger protein that resides in the nucleus. Functional analysis indicated that overexpression of NbZnFP1 and its rice orthologues triggered cell death in rice protoplasts. A NbZnFP1 orthologue was also identified in tomato and was specifically activated by AvrXa10. The results demonstrate that NbZnFP1 is a nonhost R gene that traps AvrXa10 for plant immunity in Nb.

|
Scooped by
dromius
|
(via D. Horvath, thx) Bhardwaj & Nain 2021 This review provides significant insights into the pros and cons of the two most popular genome editing tools TALENs and CRISPRs. This mini review suggests that, TALENs provides novel opportunities in the field of therapeutics being highly specific and sensitive toward DNA modifications. In this article, we will briefly explore the special features of TALENs that makes this tool indispensable in the field of synthetic biology. This mini review provides great perspective in providing true guidance to the researchers working in the field of trait improvement via genome editing.

|
Scooped by
dromius
|
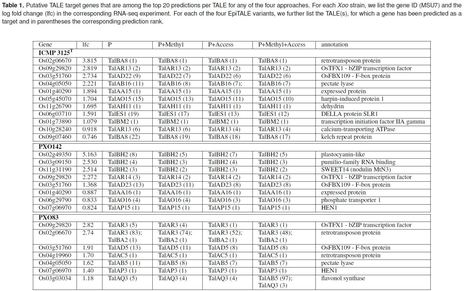
(via T. Schreiber, thx) Erkes etl al 2021 (Preprint) The yield of many crop plants can be substantially reduced by plant-pathogenic Xanthomonas bacteria. The infection strategy of many Xanthomonas strains is based on transcription activator-like effectors (TALEs), which are secreted into the host cells and act as transcriptional activators of plant genes that are beneficial for the bacteria. The modular DNA binding domain of TALEs contains tandem repeats, each comprising two hyper-variable amino acids. These repeat-variable diresidues (RVDs) bind to a continuous DNA stretch (a target box) and determine the specificity of a TALE. All available tools for the prediction of TALE targets within the host plant suffer from many false positives. In this paper we propose a strategy to improve prediction accuracy by considering the epigenetic state of the host plant genome in the region of the target box. To this end, we extend our previously published tool PrediTALE by two epigenetic features: (i) We allow for filtering target boxes according to chromatin accessibility and (ii) we allow for considering the methylation state of cytosines within the target box during prediction, since DNA methylation may affect the binding specificity of RVDs. Here, we determine the epigenetic features from publicly available DNase-seq, ATAC-seq, and WGBS-seq data in rice. We benchmark the utility of both epigenetic features separately and in combination, deriving ground-truth from RNA-seq infections studies in rice. We find an improvement for each individual epigenetic feature, but especially the combination of both. Having established an advantage in TALE target predicting considering epigenetic features, we use these data for promoterome and genome-wide scans by our new tool EpiTALE, leading to several novel putative virulence targets. Our results suggest that it would be worthwhile to collect condition-specific chromatin accessibility data and methylation information when studying putative virulence targets of Xanthomonas TALEs.

|
Scooped by
dromius
|
Becker & Boch, 2021
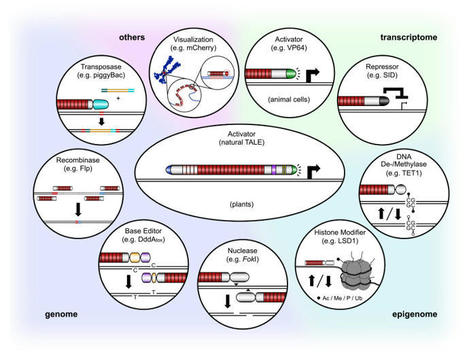
TALEN were the first easy-to-use genome editing technology and sparked the genome editing revolution. Their application in multiple species brought targeted mutagenesis to the attention of scientists worldwide. Key breakthrough successes of genome editing have since been achieved using TALEN, among these, the first commercialization of an edited crop and the first human cured from cancer. TALEN have since been largely replaced by the CRISPR technologies which are somewhat easier to build, much easier to multiplex, and have spawned multiple derived techniques. Nevertheless, the flexible and precise positioning of TALEN is unmatched, and thus they have continued to evolve to new functionalities. Here, we assemble essential facts, design guidelines as well as important past and exciting novel developments.

|
Scooped by
dromius
|
(via T. Schreiber, thx) Jain et al, 2021
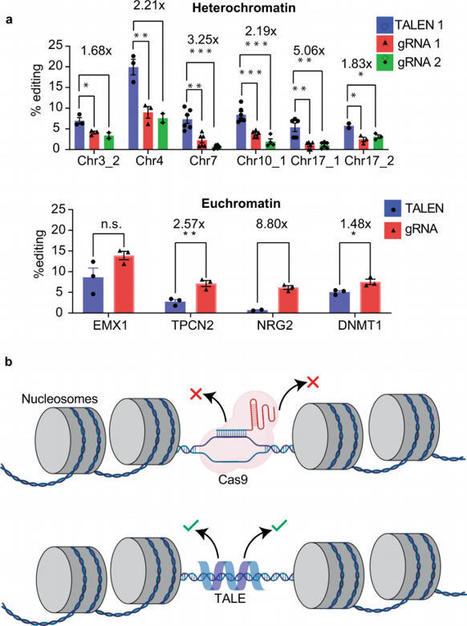
Genome editing critically relies on selective recognition of target sites. However, despite recent progress, the underlying search mechanism of genome-editing proteins is not fully understood in the context of cellular chromatin environments. Here, we use single-molecule imaging in live cells to directly study the behavior of CRISPR/Cas9 and TALEN. Our single-molecule imaging of genome-editing proteins reveals that Cas9 is less efficient in heterochromatin than TALEN because Cas9 becomes encumbered by local searches on non-specific sites in these regions. We find up to a fivefold increase in editing efficiency for TALEN compared to Cas9 in heterochromatin regions. Overall, our results show that Cas9 and TALEN use a combination of 3-D and local searches to identify target sites, and the nanoscopic granularity of local search determines the editing outcomes of the genome-editing proteins. Taken together, our results suggest that TALEN is a more efficient gene-editing tool than Cas9 for applications in heterochromatin. While Cas9 outperforms TALENs in euchromatin, it is less efficient in heterochromatic regions. Here the authors, using single-molecule imaging, show that Cas9 uses a less efficient search strategy compared to TALENs in these regions.

|
Scooped by
dromius
|
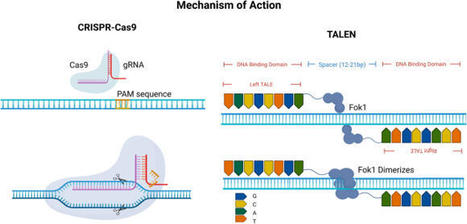
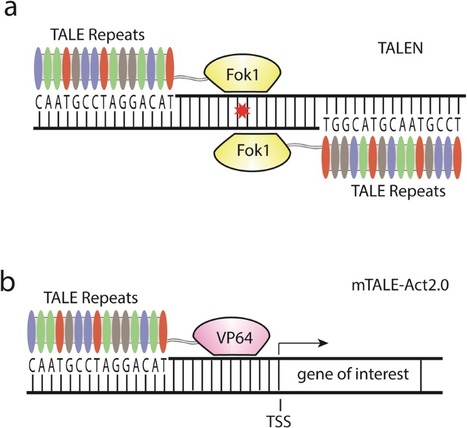
Mahlzahn & Qi 2020 Transcription activator-like effector (TALE) is a DNA-binding domain that can be paired with a nuclease to create DNA double-strand breaks, or with an effector protein to alter gene transcription. The ability to precisely alter plant genomes and transcriptomes has provided many insights into gene function and has recently been utilized for crop improvement. Easy design and construction of TALE make the tool more accessible to a variety of researchers. Here, we describe two TALE-based systems: transcription activator-like effector nucleases (TALEN), for creating targeted mutations in a gene of interest, and multiplex TALE activation (mTALE-Act), for activating one or a few genes of interest at the transcription level. Assembly of these tools is based on Golden Gate cloning and Gateway recombination, which are cost-effective and streamlined cloning methods.

|
Scooped by
dromius
|
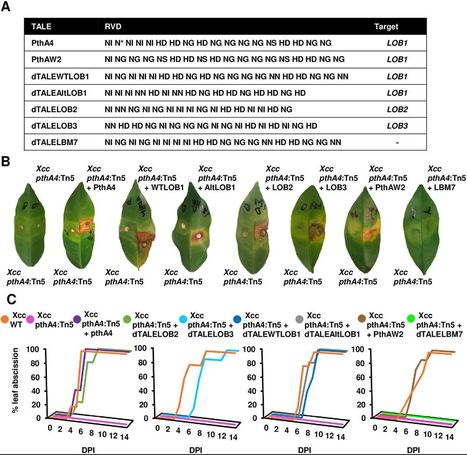
(via T. Schreiber, thx) Teper et al, 2020 Citrus canker caused by Xanthomonas citri subsp. citri (Xcc) is one of the most devastating diseases in citrus. Meiwa kumquat (Fortunella crassifolia) has shown a durable resistance against Xcc. Here, we aimed to characterize the mechanisms responsible for such a durable resistance by characterizing the transcriptional and physiological responses of Meiwa kumquat to Xcc. Inoculation of Meiwa kumquat with Xcc promoted immune responses such as upregulation of PR genes, accumulation of salicylic acid, hypersensitive response (HR)-like cell death and early leaf abscission. Hypertrophy and hyperplasia symptoms, which are known to be caused by Xcc-induction of the canker susceptibility gene LOB1 through the transcription activator-like effector (TALE) PthA4, always appear prior to the development of cell death. Mutation of pthA4 in Xcc abolished the induction of LOB1, canker symptoms, cell death, and leaf abscission and reduced the expression of PR genes in inoculated kumquat leaves without reducing Xcc titers in planta. Transcriptome analysis demonstrated that PthA4 promotes plant biotic and abiotic stress responses and the biosynthesis of abscisic acid. Transcriptional induction of LOB1 homologs in Meiwa kumquat by Xcc pthA4 mutant strains carrying a repertoire of designer TALEs promoted the elicitation of HR-like phenotype and leaf abscission, suggesting that kumquat response to Xcc is associated with upregulation of LOB1. Our study suggests a novel mechanism of plant resistance to Xanthomonas via elicitation of immune responses by upregulation of a host susceptibility gene.

|
Scooped by
dromius
|
Buchmueller et al, 2020
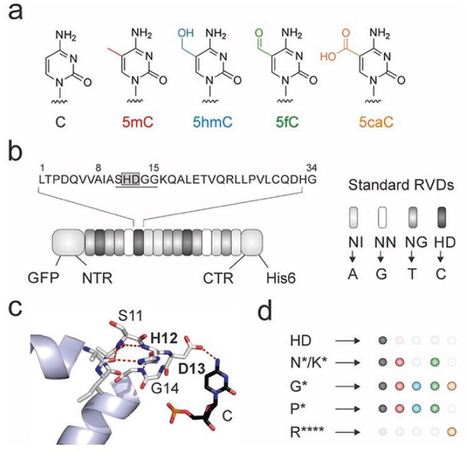
Transcription-activator like effectors (TALEs) are DNA-binding proteins used for genome targeting. TALEs contain a central domain of concatenated repeats, of which each selectively recognizes one nucleobase at the DNA major groove. Based on this simple and predictable interaction with little context dependence, TALEs offer programmable targeting of user-defined DNA sequences. Since many epigenetic DNA modifications protrude into the DNA major groove, natural and engineered TALE repeats can provide “epigenetic” selectivity, making TALEs a flexible platform to design probes for the analysis of epigenetic DNA modifications. Here, we describe guidelines for the design of TALE proteins with selectivity for epigenetic cytosine 5-modifications, the validation of their interaction with modified DNA nucleobases, and their employment in affinity enrichment assays. These techniques enable quantification of epigenetic nucleobases in user-defined genomic DNA sequences with nucleotide and strand resolution.
|

|
Scooped by
dromius
|
(via T. Lahaye, thx) Xu et al, 2022 - This study, for the first time, uncover a naturally-emerging Xa23-breaking Xoo isolate.
- The ability of AvrXa23 to be trapped by the EBE of Xa23 gene can be altered by one or more RVDs of AvrXa23.
- Seven AvrXa23-like TALEs determine the “arms-race” in the AvrXa23 and Xa23 pair.
- This study provide new insights into the diversified strategies used by Xoo to evade host resistance.
- Planting single R gene (like Xa23) rice in a great region is risk to make Xoo minority to majority.

|
Scooped by
dromius
|
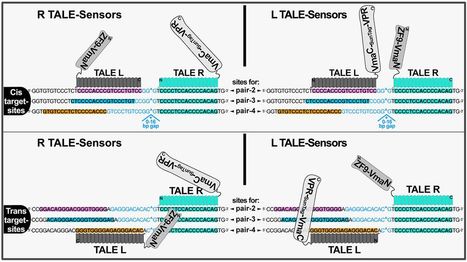
(via T. Schreiber, thx) Taghbalout et al, 2021 Here we describe TALE.Sense, a versatile platform for sensing DNA sequences in live mammalian cells enabling programmable generation of a customable response that discerns cells containing specified sequence targets. The platform is based on the programmable DNA binding of transcription activator-like effector (TALE) coupled to conditional intein-reconstitution producing a trans-spliced ON-switch for a response circuit. TALE.Sense shows higher efficiency and dynamic range when compared to the reported zinc-finger based DNA-sensor in detecting same DNA sequences. Swapping transcriptional activation modules and introducing SunTag-based amplification loops to TALE.Sense circuits augment detection efficiency of the DNA sensor. The TALE.Sense platform shows versatility when applied to a range of target sites, indicating its suitability for applications to identify live cell variants with anticipated DNA sequences. TALE.Sense could be integrated with other cellular or synthetic circuits by using specified DNA sequences as control-switches, thus expanding the scope in connecting inducible modules for synthetic biology.

|
Scooped by
dromius
|
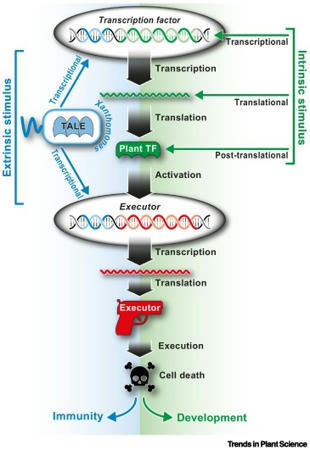
Nowak et al, 2021 Xanthomonads inject transcription-activator like effectors (TALEs) into plant cells, where TALEs bind to effector binding elements (EBEs) in plant promoters and transcriptionally activate downstream host susceptibility genes to promote disease. Some plant genotypes contain transcriptionally activatable cell death-inducing genes, termed executors, that are preceded by TALE-compatible EBEs. Such EBE-equipped executor alleles are, by definition, resistance ( R) genes, since their presence provides protection against TALE-containing xanthomonads. Recent studies uncovered that TALEs can rapidly change their sequence specificity by rearranging their modular DNA domain to circumvent detection by matching executor-type R genes. Evolutionary evidence suggests that the native function of executors is likely to be not in plant immunity, but rather in the regulation of programmed cell death, possibly in the context of plant developmental processes.

|
Scooped by
dromius
|
Haq et al, 2021 Xanthomonas oryzae pv. oryzae (Xoo), the causal agent of bacterial leaf blight in rice, delivers transcription activator-like effector (TALE) proteins into host cells to activate susceptibility or resistance (R) genes that promote disease or immunity, respectively. Nonhost plants serve as potential reservoirs of R genes; consequently, nonhost R genes may trap TALEs to trigger an immune response. In this study, we screened 17 Xoo TALEs for their ability to induce a hypersensitive response (HR) in the nonhost plant Nicotiana benthamiana (Nb); only AvrXa10 elicited an HR when transiently expressed in Nb. The HR generated by AvrXa10 required both the central repeat region and the activation domain, suggesting a specific interaction between AvrXa10 and a potential R-like gene in nonhost plants. Evans blue staining and ion leakage measurements confirmed that the AvrXa10-triggered HR was a form of cell death, and the transient expression of AvrXa10 in Nb induced immune responses. Genes targeted by AvrXa10 in the Nb genome were identified by transcriptome profiling and prediction of effector binding sites. Using several approaches (in vivo reporter assays, electrophoretic mobility-shift assays, targeted designer TALEs, and on-spot gene silencing), we confirmed that AvrXa10 targets NbZnFP1, a C2H2-type zinc finger protein that resides in the nucleus. Functional analysis indicated that overexpression of NbZnFP1 and its rice orthologs triggered cell death in rice protoplasts. An NbZnFP1 ortholog was also identified in tomato and was specifically activated by AvrXa10. These results demonstrate that NbZnFP1 is a nonhost R gene that traps AvrXa10 to promote plant immunity in Nb.

|
Scooped by
dromius
|
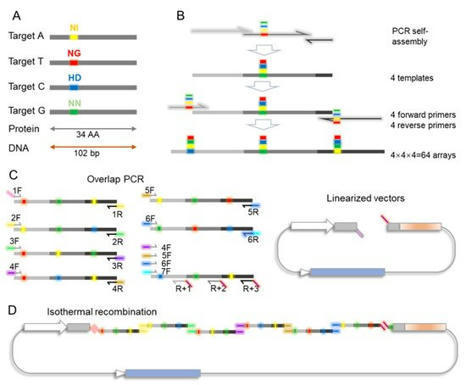
(via T. Schreiber and T. Lahaye, thanks!) Cheng et al. 2021 Transcription activator-like effectors (TALEs) have been effectively used for targeted genome editing, transcriptional regulation, epigenetic modification, and locus-specific DNA imaging. However, with the advent of the clustered regularly interspaced short palindromic repeat/Cas9 system, an easy-to-use tool with the same function as TALEs, TALEs have recently been abandoned because of their complexity, time consumption, and difficult handling in common labs. Here, we described a degenerated codon-based TALE assembly system for simple, rapid, and efficient TALE assembly. TALE trimers with nonrepetitive DNA sequences were amplified by PCR and sequentially assembled via Gibson assembly. Our method is cost-effective, requires only commonly used basic molecular biology reagents, and takes only 2 h from target sequence analysis to completion. This simple, rapid, and lab-friendly TALE assembly method will restore the value of TALEs in DNA targeting.

|
Scooped by
dromius
|
(via T. Schreiber, thx) Tse et al, 2021 Architectural proteins alter the shape of DNA. Some distort the double helix by introducing sharp kinks. This can serve to relieve strain in tightly-bent DNA structures. Here, we design and test artificial architectural proteins based on a sequence-specific Transcription Activator-like Effector (TALE) protein, either alone or fused to a eukaryotic high mobility group B (HMGB) DNA-bending domain. We hypothesized that TALE protein binding would stiffen DNA to bending and twisting, acting as an architectural protein that antagonizes the formation of small DNA loops. In contrast, fusion to an HMGB domain was hypothesized to generate a targeted DNA-bending architectural protein that facilitates DNA looping. We provide evidence from Escherichia coli Lac repressor gene regulatory loops supporting these hypotheses in living bacteria. Both data fitting to a thermodynamic DNA looping model and sophisticated molecular modeling support the interpretation of these results. We find that TALE protein binding inhibits looping by stiffening DNA to bending and twisting, while the Nhp6A domain enhances looping by bending DNA without introducing twisting flexibility. Our work illustrates artificial approaches to sculpt DNA geometry with functional consequences. Similar approaches may be applicable to tune the stability of small DNA loops in eukaryotes.

|
Scooped by
dromius
|
Yoshihisa et al. 2021
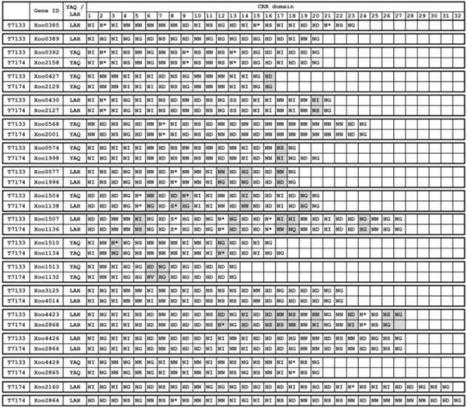
Xanthomonas oryzae pv. oryzae (Xoo) possesses transcription activator-like (TAL) effectors, which are important virulence factors for bacterial blight disease. Rice nucleotide-binding leucine rich repeats receptor Xa1 recognizes TAL effectors, then induces various immune responses. To overcome host defenses, Xoo has developed TAL effector variants, termed interfering TAL (iTAL) (or truncated TAL) effectors. Japanese Xoo strain T7133 overcomes Xa1-mediated immunity. In this study, we determined the whole genome sequence of Xoo T7133 and identified TAL and iTAL effectors, revealing commonalities and differences in the TAL effectors between Xoo T7133 and avirulent strain T7174. In addition, Xoo T7133 has a different type of iTAL effector from Xoo T7174, suggesting that the iTAL effector of Xoo T7133 may contribute to the suppression of Xa1-mediated immunity.

|
Scooped by
dromius
|
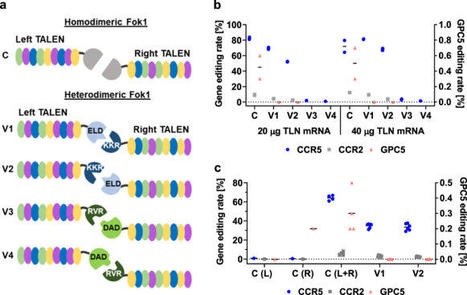
(via T. Schreiber, thx) Schwarze et al, 2021 Disruption of the C-C-Chemokine-receptor-5 (CCR5) gene induces resistance towards CCR5-tropic HIV. Here we optimised our previously described CCR5-Uco-TALEN and its delivery by mRNA electroporation. The novel variant, CCR5-Uco-hetTALEN features an obligatory heterodimeric Fok1-cleavage domain, which resulted in complete abrogation of off-target activity at previously found homodimeric as well as 7/8 in silico predicted, potential heterodimeric off-target sites, the only exception being highly homologous CCR2. Prevailing 18- and 10-bp deletions at the on-target site revealed microhomology-mediated end-joining as a major repair pathway. Notably, the CCR5Δ55–60 protein resulting from the 18-bp deletion was almost completely retained in the cytosol. Simultaneous cutting at CCR5 and CCR2 induced rearrangements, mainly 15-kb deletions between the cut sites, in up to 2% of T cells underlining the necessity to restrict TALEN expression. We optimised in vitro mRNA production and showed that CCR5-on- and CCR2 off-target activities of CCR5-Uco-hetTALEN were limited to the first 72 and 24–48 h post-mRNA electroporation, respectively. Using single-cell HRMCA, we discovered high rates of TALEN-induced biallelic gene editing of CCR5, which translated in large numbers of CCR5-negative cells resistant to HIVenv-pseudotyped lentiviral vectors. We conclude that CCR5-Uco-hetTALEN transfected by mRNA electroporation facilitates specific, high-efficiency CCR5 gene-editing (30%–56%) and it is highly suited for clinical translation subject to further characterisation of off-target effects.

|
Scooped by
dromius
|
Lee et al, 2021
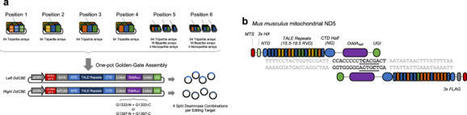
DddA-derived cytosine base editors (DdCBEs), composed of the split interbacterial toxin DddAtox, transcription activator-like effector (TALE), and uracil glycosylase inhibitor (UGI), enable targeted C-to-T base conversions in mitochondrial DNA (mtDNA). Here, we demonstrate highly efficient mtDNA editing in mouse embryos using custom-designed DdCBEs. We target the mitochondrial gene, MT-ND5 (ND5), which encodes a subunit of NADH dehydrogenase that catalyzes NADH dehydration and electron transfer to ubiquinone, to obtain several mtDNA mutations, including m.G12918A associated with human mitochondrial diseases and m.C12336T that incorporates a premature stop codon, creating mitochondrial disease models in mice and demonstrating a potential for the treatment of mitochondrial disorders.

|
Scooped by
dromius
|
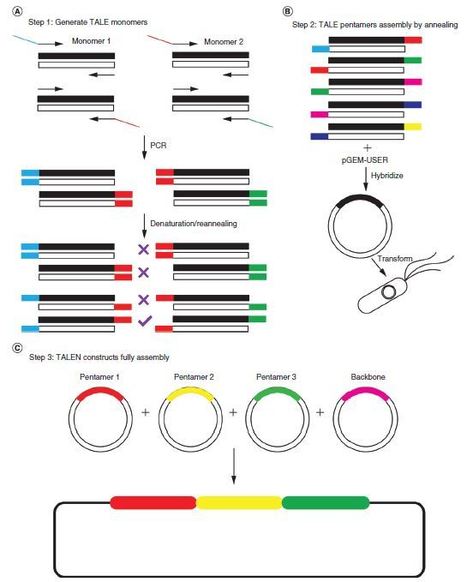
Wang et al., 2021 Transcription activator-like effector (TALE) nucleases (TALENs) efficiently recognize and cleave DNA in a sequence-dependent manner. However, current TALE custom synthesis methods are either complicated or expensive. Here we report a simple and low-cost method for TALE construct assembly. This method utilizes the denaturation/reannealing nature of double-stranded DNA to create a unique single-stranded DNA overhang for proper ordering of TALEmonomers in an engineered multimer.We successfully synthesized two TALEN pairs targeting the endogenous TET1
locus in human embryonic kidney cells and demonstrated their editing efficiency. Our method provides an alternative simple, low-cost method for effective TALEN assembly, which may improve the application of TALE-based technology.

|
Scooped by
dromius
|
Bacman & Moraes 2020
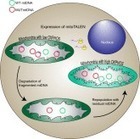
The mixture of mutant and wild-type mitochondrial DNA (mtDNA) in cells from patients with mitochondrial diseases provides an opportunity to develop therapies by selectively eliminating the mutant fraction. Our laboratory has adapted TALENs, a gene editing platform, to specifically cleave mutant mtDNA. Ex vivo and in vivo experiments have provided proof-of-principle that the approach works in changing mtDNA heteroplasmy toward the wild-type mtDNA.

|
Scooped by
dromius
|
Liu, Hubbard, Badran 2020, US Patent application Engineered transcriptional activator-like effectors (TALEs) are versatile tools for genome manipulation with applications in research and clinical contexts. One current drawback of TALEs is that the 5′ nucleotide of the target is specific for thymine (T). TALE domains with alternative 5′ nucleotide specificities could expand the scope of DNA target sequences that can be bound by TALEs. Another drawback of TALEs is their tendency to bind and cleave off-target sequence, which hampers their clinical application and renders applications requiring high-fidelity binding unfeasible. This disclosure provides methods and strategies for the continuous evolution of proteins comprising DNA-binding domains, e.g., TALE domains. In some aspects, this disclosure provides methods and strategies for evolving such proteins under positive selection for a desired DNA-binding activity and/or under negative selection against one or more undesired (e.g., off-target) DNA-binding activities. Some aspects of this disclosure provide engineered TALE domains and TALEs comprising such engineered domains, e.g., TALE nucleases (TALENs), TALE transcriptional activators, TALE transcriptional repressors, and TALE epigenetic modification enzymes, with altered 5′ nucleotide specificities of target sequences. Engineered TALEs that target ATM with greater specificity are also provided.
|



 Your new post is loading...
Your new post is loading...











https://pijnpillen.com/Producten/buy-flakka-a-pvp/
https://pijnpillen.com/Producten/buy-ultram-online/
https://pijnpillen.com/Producten/ketamine-poeder-kopen/
https://pijnpillen.com/Producten/buy-micro-mushrooms/
https://pijnpillen.com/Producten/carisoprodol-nederland-kopen/
https://pijnpillen.com/Producten/koop-natriumcyanide/
https://pijnpillen.com/Producten/koop-vicodin-online/
https://pijnpillen.com/Producten/koop-dmt-netherland/
https://pijnpillen.com/Producten/koop-dexedrine/
https://pijnpillen.com/Producten/koop-vyvanse-online/
https://pijnpillen.com/Producten/koop-morphine/
https://pijnpillen.com/Producten/buy-4-aco-dmt-usa/
https://pijnpillen.com/Producten/oxycodon-hcl-kopen/
https://pijnpillen.com/Producten/koop-suboxone-strips/
https://pijnpillen.com/Producten/koop-percocet-online/
https://pijnpillen.com/Producten/koop-oxycontin-online/
https://pijnpillen.com/Producten/buy-flakka-a-pvp/
https://pijnpillen.com/Producten/fentanyl-pleister-kopen/
https://pijnpillen.com/Producten/buy-seconal-sodium/
https://pijnpillen.com/Producten/fentanyl-poeder-kopen/
https://pijnpillen.com/Producten/koop-morphine/
https://pijnpillen.com/Producten/buprenorfine-kopen/
https://pijnpillen.com/Producten/koop-vyvanse-online/
https://pijnpillen.com/Producten/carisoprodol-nederland-kopen/
https://pijnpillen.com/Producten/koop-natriumcyanide/
https://pijnpillen.com/Producten/koop-dmt-netherland/
https://pijnpillen.com/Producten/koop-vicodin-online/
https://pijnpillen.com/Producten/gouden-leraar-paddenstoelen/
%3C/a""> ">buy-flakka-a-pvp
buy-ultram-online
%3C/a""> ">ketamine-poeder-kopen
%3C/a""> ">buy-micro-mushrooms
carisoprodol-nederland-kopen
koop-natriumcyanide
%3C/a""> ">koop-vicodin-online
%3C/a""> ">koop-dmt-netherland
koop-dexedrine
%3C/a""> ">koop-morphine
buy-4-aco-dmt-usa
oxycodon-hcl-kopen
koop-suboxone-strips
%3C/a""> ">koop-percocet-online
acheter adipex
buy-flakka-a-pvp
%3C/a""> ">fentanyl-pleister-kopen
buy-seconal-sodium
fentanyl-poeder-kopen
koop-morphine
buprenorfine-kopen
koop-vyvanse-online
carisoprodol-nederland-kopen
koop-natriumcyanide
koop-dmt-netherland
koop-vicodin-online
gouden-leraar-paddenstoelen
https://acheteroxycodone.eu/produit/comprar-cianuro-de-sodio/